Tianjin/FLIP-FLOP
From 2007.igem.org
DesignFlip-Flop, a huge family of basic electric elements, is widely applied to the field of electric circuit and construction of database. One of the most common members of this family is RS Flip-Flop, the key part of which is a clock-controlled element. Based on the the “Latch and Enable Control" conception of "Flip-Flop" and synthetic biology, we designed the Genetical RS Flip-Flop whose output signal (Green Fluorescence protein) is regulated by input signal (the addition of IPTG). Besides, we modulate the performance of Genetical RS FLIP-FLOP to optimize our original design. 1.Introduction to the Logic Rules of Our Flip-Flop
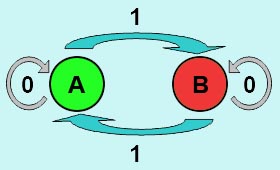
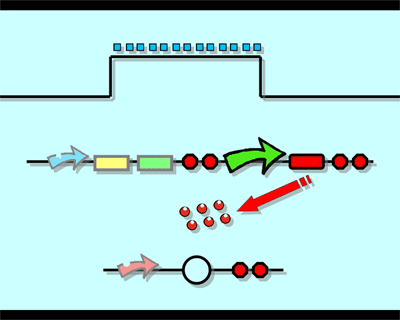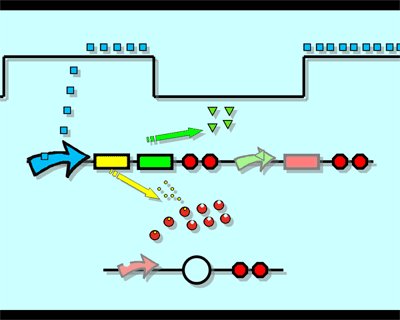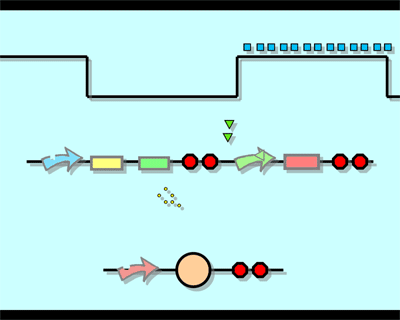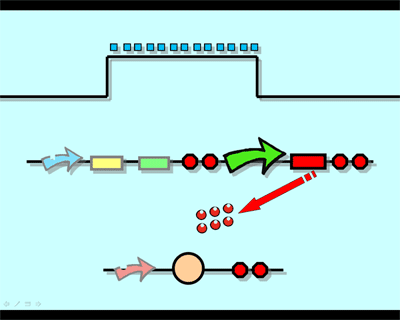
The logic principle of our design is shown above. Unless the input signal transfer from one stable condition to another (such as 0 to 1 or 1 to 0), the output signal would change into 1, otherwise it would maintain 0. Thus, the immediate response to emergency (input change) enables our design to detect signal variation in a short time, which is beneficial to the control of biological systems.
| ||||||
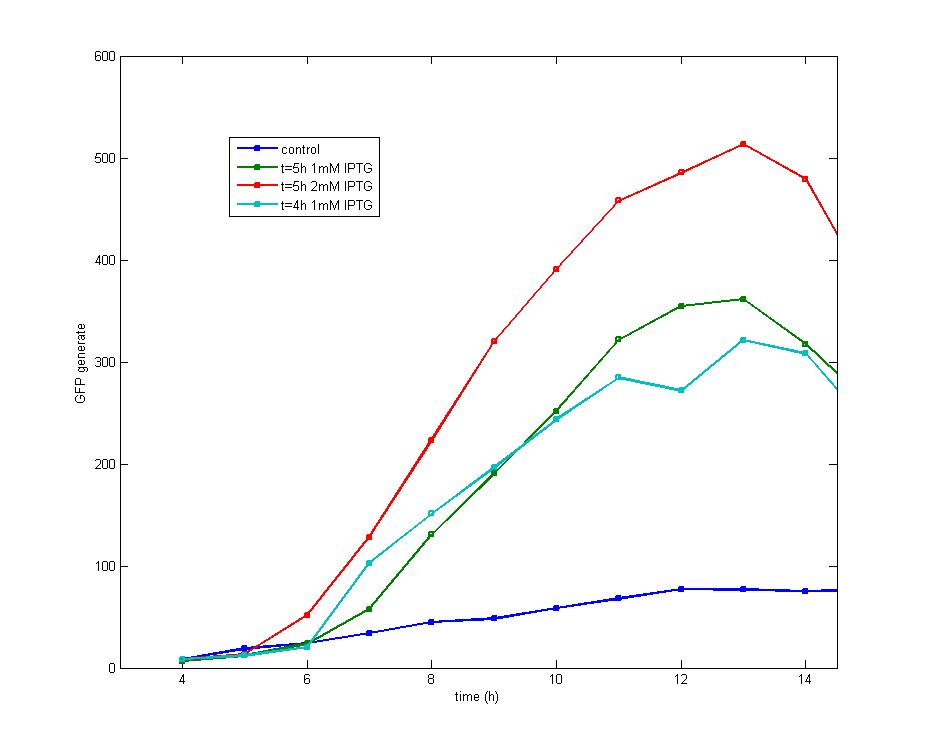
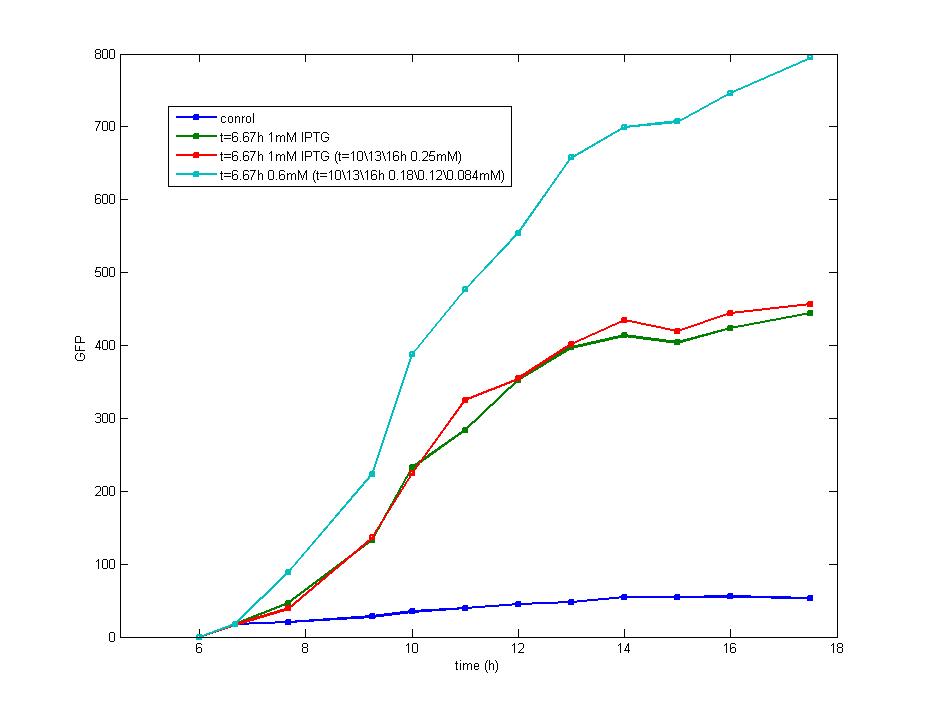
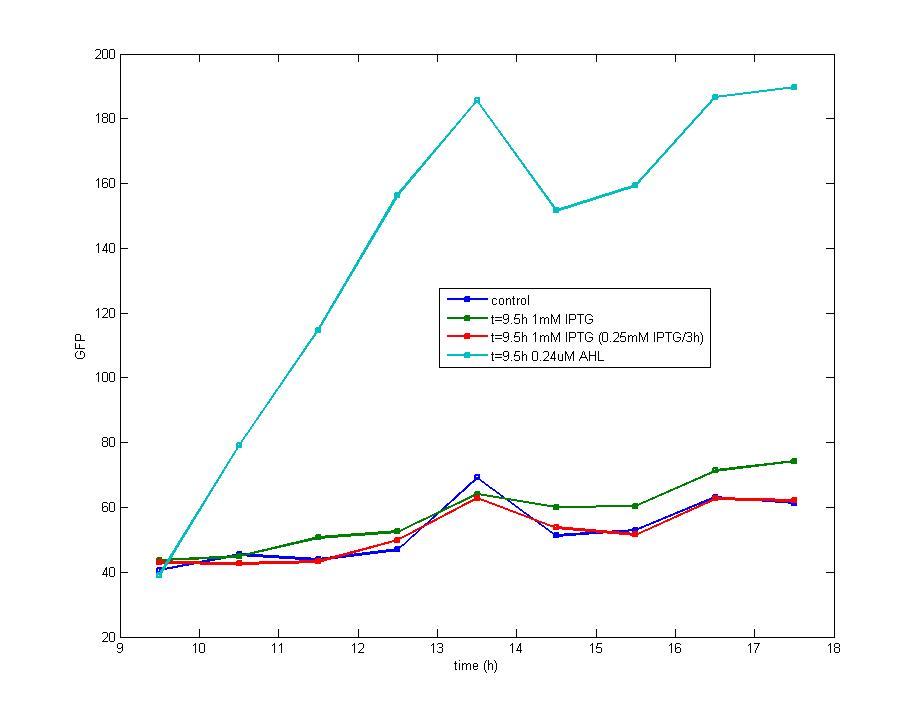
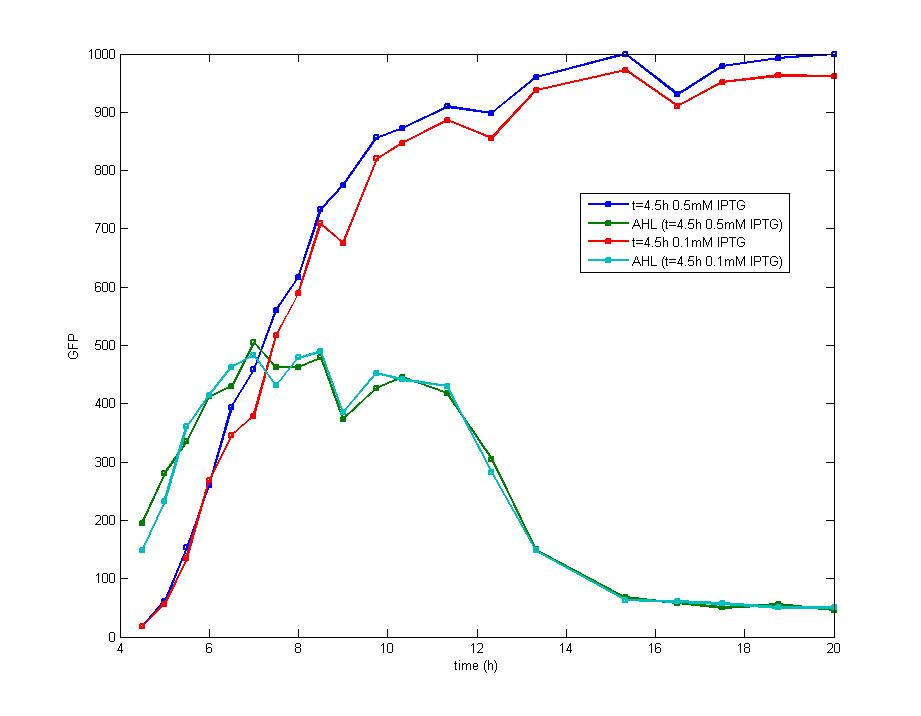
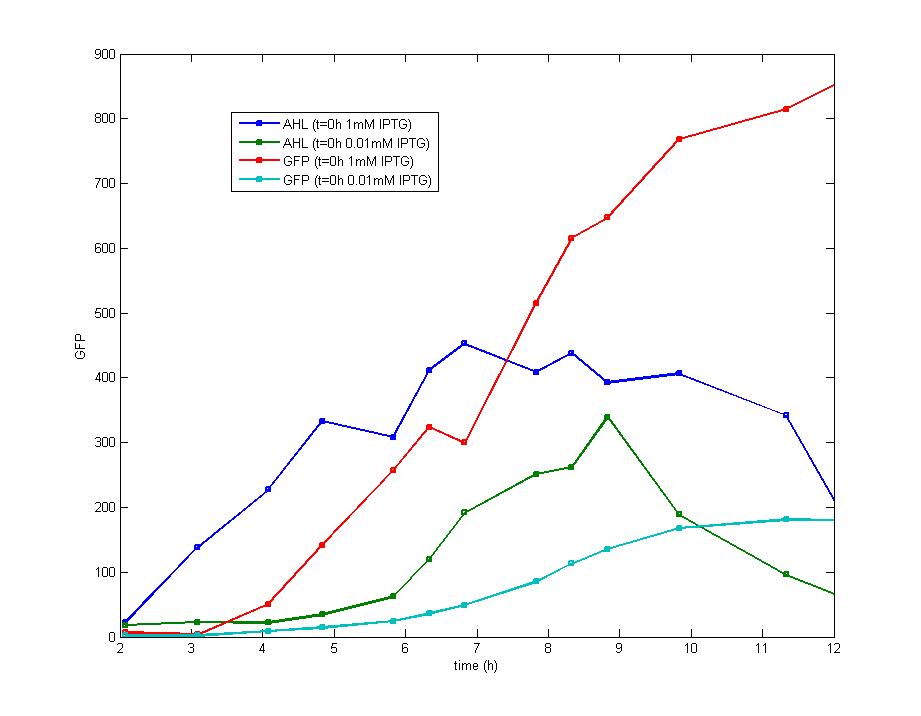
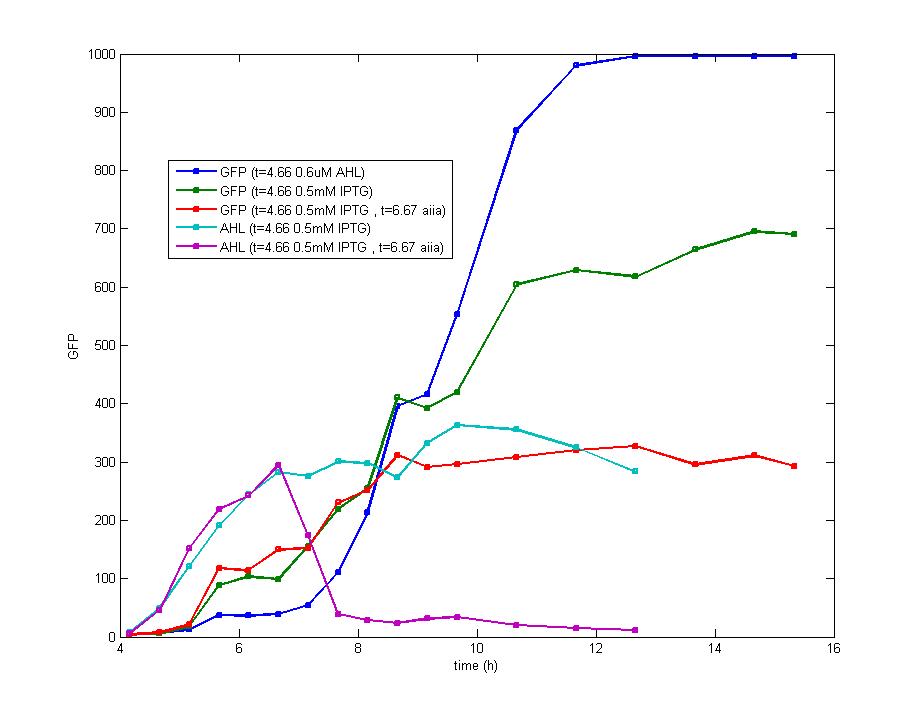
ModelingMainly based on Ordinary Differential Equations, a mathematical model of our Flip-Flop system is constructed to test the result of our design and predict potential key factors deciding the results of our experiment. According to the model, the variation of output signal responding to the input signal matches the typical feature of Flip-Flop, and there would be no output signal unless at the positive edge and negative edge of the input signal. The influence of AHL to the output signal is also considered. By changing the degradation rate of AHL, the fluorescence intensity exhibits different behaviour which could be explained by principles of Flip-Flop. Finally, the parameter sensitivity is also tested to explore most significant parameters to output signals and it is discovered that the strength of promoter I, which controls the production of LuxI, the promoter II, which controls the expression of LuxR, exert great effect on the final results. 1. Construction of Mathematical Model
| ||||||
Experiment2.
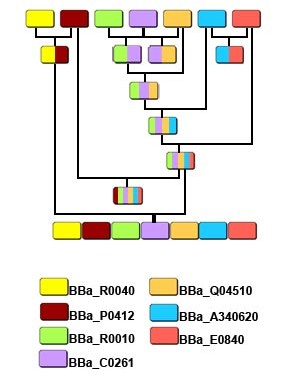
Parts Reservoir
|