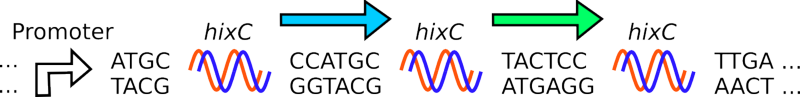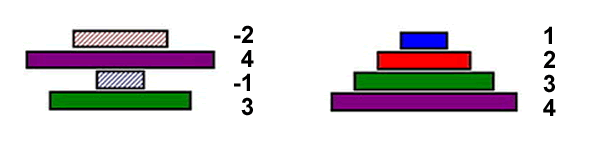Davidson Missouri W/Background Information
From 2007.igem.org
| Line 1: | Line 1: | ||
| - | <center>[[Davidson Missouri W| <span style="color:red">Home</span>]] | [[Davidson Missouri W/Background Information| <span style="color:black">Background Information</span>]] | [[Davidson Missouri W/Solving the HPP in vivo| <span style="color:red">Current Project: Solving the Hamiltonian Path Problem ''in vivo''</span>]] | [[Davidson Missouri W/Mathematical Modeling| <span style="color:red">Mathematical Modeling</span>]] | [[Davidson Missouri W/Gene splitting| <span style="color:red"> Gene Splitting </span>]] | [[Davidson Missouri W/Controlling Expression| <span style="color:red"> Controlling Expression </span>]] | [[Davidson Missouri W/Traveling Salesperson Problem| <span style="color:red">Traveling Salesperson Problem | + | <center>[[Davidson Missouri W| <span style="color:red">Home</span>]] | [[Davidson Missouri W/Background Information| <span style="color:black">Background Information</span>]] | [[Davidson Missouri W/Solving the HPP in vivo| <span style="color:red">Current Project: Solving the Hamiltonian Path Problem ''in vivo''</span>]] | [[Davidson Missouri W/Mathematical Modeling| <span style="color:red">Mathematical Modeling</span>]] | [[Davidson Missouri W/Gene splitting| <span style="color:red"> Gene Splitting </span>]] | [[Davidson Missouri W/Controlling Expression| <span style="color:red"> Controlling Expression </span>]] | [[Davidson Missouri W/Traveling Salesperson Problem| <span style="color:red">Traveling Salesperson Problem</span> ]] | [[Davidson Missouri W/Resources and Citations|<span style="color:red">Resources and Citations</span>]]</center> |
<hr> | <hr> | ||
Revision as of 20:49, 26 September 2007
In 1994 Adleman developed a system to solve the Hamiltonian Path problem using DNA in vitro. We have implemented a bacterial system to solve Hamiltonian Path problems in vivo. Our bacterial computer works by taking advantage of the Hin Recombinase protein and HixC sites to perform a "flipping" operation. By strategically placing HixC sites on a plasmid, along with reporter genes, we can simulate paths on a graph through flipping.
The Hin Recombinase/HixC System Revisited
Salmonella typhimurium is a bacterium which moves using flagella. Depending on its environment, it expresses different sets of surface proteins on its flagella. It is able to change this expression through the use of Hin Recombinase and HixC sites. The Hin Recombinase enzyme catalyzes the inversion of DNA that lies between a pair of HixC sites. By flipping DNA segments between HixC sites S. typhimurium can express alternate flagellar surface proteins.
- Given a segment of DNA, flanked by HixC sites (blue arrow):
- Two dimers of Hin Recombinase protein attach to a pair of HixC sites around the DNA segment.
- The DNA segment between the pair of HixC sites is inverted. Note that the green arrow is unaffected.
Flipping Pancakes
In the 2006 iGEM competition, Davidson and Missouri Western used Hix Recombinase and HixC to model a mathematical problem, called the Pancake Flipping problem. The problem is the following: given a stack of pancakes which are burnt on one side and golden on the other, and two spatulas, what is the minimum number of flips required to sort the stack by pancake size and with all burnt sides facing downwards?
Using mathematical models it is possible to compute these estimates for small stacks. However, adding pancakes to a stack increases the number of possible permutations non-linearly. The number of possible stacks of n pancakes is equal to: 2^n (n!). Thus traditional computers are not well-suited for computations involving large pancake stacks.
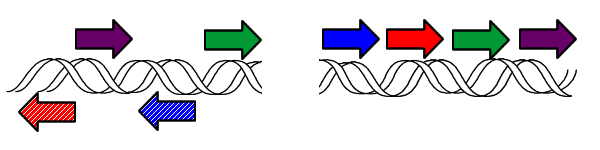
However, it is possible to model pancake flipping using bacteria. If each pancake is given an identification number, along with a sign designating its orientation, then this can be stated equivalently using genes. Thus the following images show equivalent notations for pancake stacks:
By taking advantage of this notation it is possible to use bacteria as computers to solve pancake-flipping problems.
Why Use Bacteria?
Although we have demonstrated how it is possible to solve mathematical problems using bacteria, it may not be obvious why this is helpful. Due to the exponential increase in problem size that arises from a linear increase in pancake stack size, traditional computers are not good at solving large pancake problems. However, bacterial computers are not affected as much by this problem. If each bacterium contains several plasmids, each plasmid is a computer, and a single flask of bacteria contains millions of bacteria, then it is trivial to produce millions of computers. Each computer will be acting in parallel with the others, drastically reducing the amount of time required to reach a solution. The use of bacterial computers is thus a way to allow us to solve computational problems which traditional computers cannot.