Glasgow/Wetlab/Resources
From 2007.igem.org
(→Plasmids) |
|||
| Line 62: | Line 62: | ||
!width="180" |<font face=georgia color=#3366CC size=4>pQF52</font> | !width="180" |<font face=georgia color=#3366CC size=4>pQF52</font> | ||
<br> [[Glasgow/Wetlab/References|Tran ''et al'', 1998]] | <br> [[Glasgow/Wetlab/References|Tran ''et al'', 1998]] | ||
| - | || [[Image:PQF52.PNG]] | + | || [[Image:PQF52 2.PNG]] |
|} | |} | ||
Latest revision as of 15:21, 26 October 2007
 | Back To Glasgow's Main Page | Back To Glasgow's Wetlab Log |
|---|
Resources
Some resources we have found useful in our research.
| BioEdit | Biological sequence alignment editor |
|---|---|
| BLAST | Compares nucleotide or protein sequences to sequence databases and calculates the statistical significance of matches. |
| NCBI | National Center fot Biotechnology Information. |
| Open Wetware | DNA sequence analysis. |
| pDRAW32 | DNA sequence analysis. |
| Primer3 | Checking primer design. |
| Primer Melting Temperatures | Calculating primer melting temperatures for DNA sequencing. |
| Webcutter 2.0 | Restriction mapping. |
Plasmids
Diagrams of the sequences from which we made our parts.
| XylR, pu and pr inpGLTUR | 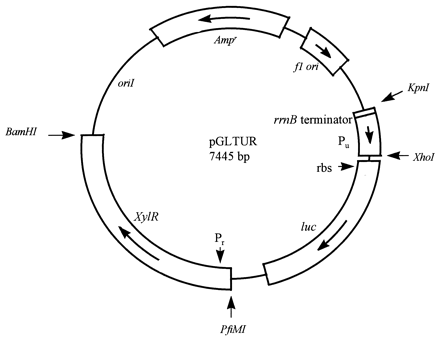
|
|---|---|
| DntR and pDntR | 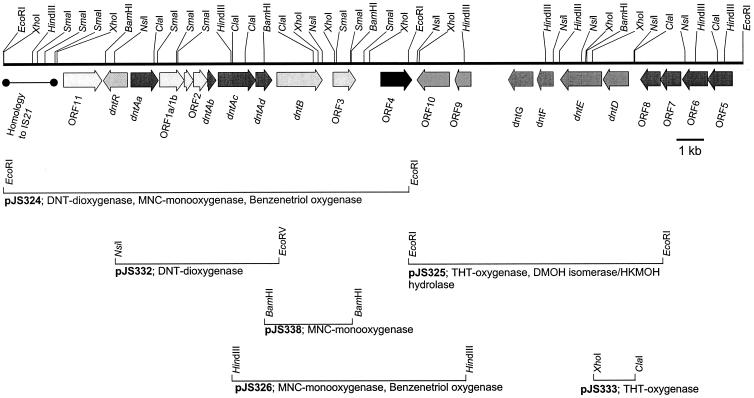
|
| pQF52 | 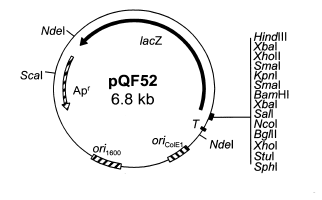
|
 | Back To Glasgow's Main Page | Back To Glasgow's Wetlab Log |
|---|