Paris/July 16
From 2007.igem.org
Contents |
Plasmid pKs::DGAT expression in E. Coli
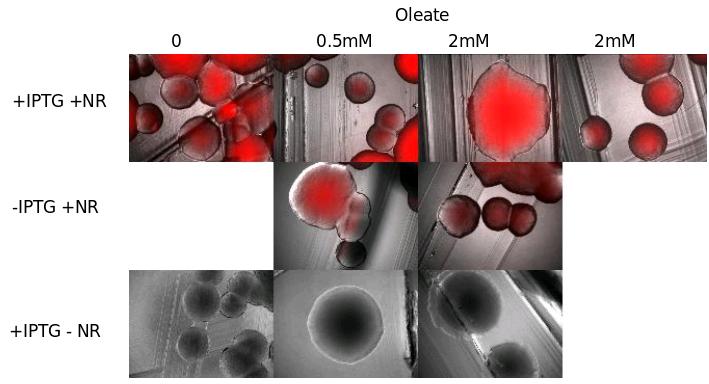
We tried to see TG with NR died on E. Coli cells transfected with pKs::DGAT, an IPTG-inducible promoter (see July 14) . We tried different growth media containing more or less oleate (0.5 and 2mM), that should in theory increase TG synthesis.
- Observations:
- Interpretation:
We don't see the inducible effect of IPTG.
We can think that :
- Either the fluorescence without IPTG is due to a leak of the promoter.
- Either DGAT is not induce in presence of IPTG, and the fluorescence we see is only a background.
- Perspectives:
- To explain the lack of the promoter pLac, we can notice that we are here in stationary phase, the bacterium metabolism can also change and the presence of lactose derepress LacI leading to expression of DGAT without adding IPTG in the medium.
We also decided to test synthesis of TG in exponential phase.
- To see if the fluorescence is only background, we can use as control E.coli non-transformed by pKS::DGAT.
- We can also wait to see accumulation of TG. Indeed in Acinetobacter, the process is pretty long (2-4 days). It could be the same for E.coli transformed by pKS::DGAT.
We decided to incubate E.coli DGAT and non transformed by pKS::DGAT (transformed by part B0015, plasmid with ampiR) in LB ampicillin medium overnight.
Growth kinetics of w121 strain
Results of the previous day : we lost everything because of a crash of the computer :( Sorry Eimad !
Transduction of MG1655 with P1 stock made on w121
- Control (1mL LB MgSO4 30mM; CaCl2 15mM)
- 5µL Phage + 900µL LB (MgSO4 30mM; CaCl2 15mM) + 100µL MG1655 Culture ON
- 50µL Phage + 900µL LB (MgSO4 30mM; CaCl2 15mM) + 100µL MG1655 Culture ON
- 500µL Phage + 500µL LB (MgSO4 30mM; CaCl2 15mM) + 100µL MG1655 Culture ON
=>For the nth time, it is not working : we have only contaminants.
MiniPreps
- I0500 clones 1, 2
- pJ23107 clones 1, 2
PCR purification
- Lox71-FtsZ1
- FtsZ2
- DGAT1
- DGAT2
PCRs
The Lox66-DapAColi PCR did not work... We'll try again with different annealing temperature
Assembly PCRs:
- Lox71-FtsA-FtsZ-1 + FtsZ-2
- DGAT-1 + DGAT-2
The Lox66-DapAColi PCR did not work... so we try a gradient of annealing temperatures
Transformations of Biobricks
As in the protocol given in the registry. Transformation of biobricks :
- BBa_B0030 pSB1A2 : RBS (well 3G plate 1)
- BBa_E0422 pSB1A2 : ECFP (RBS+LVA+Term) (well 11G plate 1)
- BBa_E0241 pSB1A2 : PoPs to GFP converter (well 15c plate 2)
- BBa_E0840 gfp tri-part; strong rbs (well 16E plate 1)
- BBa_J61047 pSB1A2 : Cre ORF (well 8P plate 4)
Transformation in DH5alpha subcloning efficiency
Spread on LB-Amp