Splitting Genes for HPP
From 2007.igem.org
Macampbell (Talk | contribs) |
Macampbell (Talk | contribs) (→'''Splitting Reporter Genes with HixC''') |
||
| Line 1: | Line 1: | ||
| - | + | # Red Fluorescent Protein | |
| - | + | # Kanamycin Nucleotidyltransferase | |
| + | # Cre Recombinase | ||
| + | # Chloramphenicol Acetyl Transferase | ||
| + | <br> | ||
| + | ---- | ||
'''DsRed - Red Fluorescent Protein''' | '''DsRed - Red Fluorescent Protein''' | ||
| Line 8: | Line 12: | ||
[[Image:Rfp hix insertion point.jpg]] | [[Image:Rfp hix insertion point.jpg]] | ||
| + | <br> | ||
| + | ---- | ||
'''Kanamycin Nucleotidyltransferase''' | '''Kanamycin Nucleotidyltransferase''' | ||
| Line 39: | Line 45: | ||
We decided to insert our hix sites at the 125 AA of each monomer due to their distance from each other, active site secondary structure, N or C terminus, and lack of any previous mutational analysis proving its function as integral. | We decided to insert our hix sites at the 125 AA of each monomer due to their distance from each other, active site secondary structure, N or C terminus, and lack of any previous mutational analysis proving its function as integral. | ||
| + | <br> | ||
| + | ---- | ||
'''Chloramphenicol Acetyltransferase''' | '''Chloramphenicol Acetyltransferase''' | ||
| + | <br> | ||
| + | ---- | ||
'''Cre Recombinase''' | '''Cre Recombinase''' | ||
| Line 49: | Line 59: | ||
[http://www3.interscience.wiley.com/cgi-bin/abstract/104558885/ABSTRACT?CRETRY=1&SRETRY=0]. | [http://www3.interscience.wiley.com/cgi-bin/abstract/104558885/ABSTRACT?CRETRY=1&SRETRY=0]. | ||
| + | |||
| + | <br> | ||
| + | ---- | ||
Revision as of 18:17, 14 June 2007
- Red Fluorescent Protein
- Kanamycin Nucleotidyltransferase
- Cre Recombinase
- Chloramphenicol Acetyl Transferase
DsRed - Red Fluorescent Protein
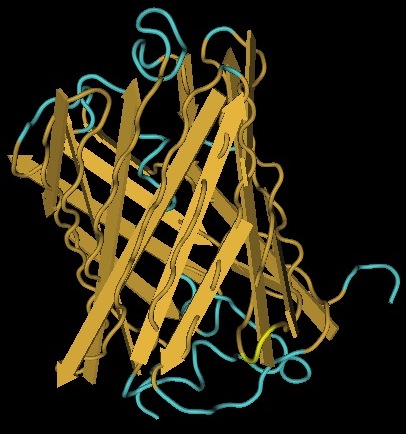
We use genes to represent the nodes on our Hamiltonian path. One of the essential features of these genes is that they can tolerate the insertion of a Hix site. It has been previously demonstrated that GFP fluoresces despite a Hix insertion. Another glowing protein, [http://partsregistry.org/Part:BBa_E1010 RFP] (from [http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?id=86600 Discosoma sp.]), is a candidate for use in our path. Although its DNA sequence is markedly different from GFP's, it has some amino acid similarity and a remarkable structural similarity. Both proteins have a Beta-barrel structure which surrounds an internal chromophore.
Inserting 13 amino acids into a protein can potentially disrupt its ability to function. It is thus essential to find an insertion point that does not interfere with the protein's function. Fortunately, the similarity between GFP and RFP allows us to make a highly educated guess for where to insert. RFP's amino acid position 154 is homologous to GFP's amino acid position 157, which is where GFP was split. This is therefore our best guess for where to insert the Hix site.
Kanamycin Nucleotidyltransferase
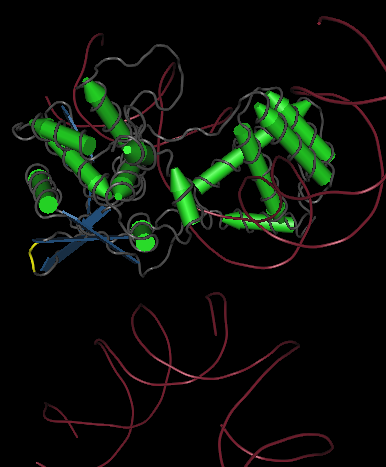
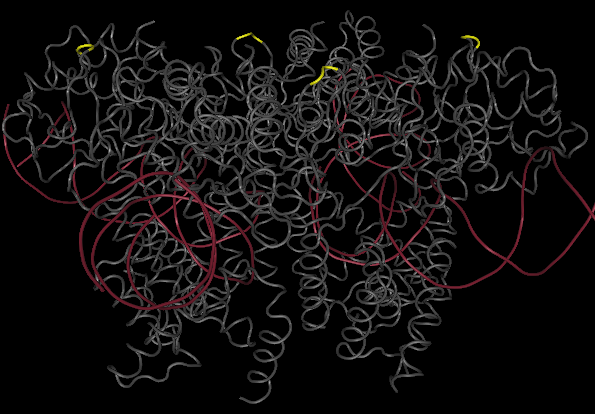
One gene our team will be using as a node in our Hamiltonian Path problem is Kanamycin resistance translated in the form of Kanamycin nucleotidyltransferase (KNTase). The antibiotic Kanamycin, once in the cytosol of E.Coli, inhibits protein synthesis by interacting with the “decoding” region in the small ribosomal subunit RNA.(Sambrook and Russel, 2001) The KNTase enzyme, as a member of the aminoglycoside phosphotransferase (APH) enzyme family, blocks Kanamycin’s ability to inhibit protein synthesis by transferring a nucleoside monophosphate (adenyl) group from Mg2+-ATP to the 4’ hydroxyl group of Kanamycin, inhibiting its ability to bind to the srRNA.[http://www.ingentaconnect.com/content/els/00452068/1999/00000027/00000005/art91144 1] Our goal was to insert a hix site (a polar molecule) in an area of KNTase protein that would not interfere with its ability to inhibit Kanamycin. We looked at mutational analysis of KNTase and other aminoglycoside phosphotransferase enzymes to determine which aspects of KNTase’s structure were integral to its function and therefore not an ideal site for hix site insertion. KNTase is a dimmer consisting of 253 amino acids in the molecule [http://www.bioscience.org/1999/v4/d/wright/fulltext.htm 3]. In looking at conserved structures in the APH family we took into consideration that:
-Substitution of AA 190 caused 650-fold decrease in enzyme activity [http://www.bioscience.org/1999/v4/d/perlin/fulltext.htm 2]
-AA 190 is involved in catalysis [http://www.bioscience.org/1999/v4/d/wright/fulltext.htm 5]
-AA 195 and 208 are involved in Mg2+ binding [http://www.bioscience.org/1999/v4/d/wright/fulltext.htmv 5]
-Mutant Enzymes 190, 205, 210 all showed changes in mg+2 binding from the WT [http://www.bioscience.org/1999/v4/d/perlin/fulltext.htm 2]
-Substitution of AA 210 (conserved) reduced enzyme activity [http://www.bioscience.org/1999/v4/d/perlin/fulltext.htm 2]
-AA 166 serves to catalyze reactions involving ATP [http://www.bioscience.org/1999/v4/d/perlin/fulltext.htm 2]
-AA 44 is involved in ATP binding [http://www.bioscience.org/1999/v4/d/wright/fulltext.htm 5]
-AA 60 is involved in orientation of AA 44 and ATP binding [http://www.bioscience.org/1999/v4/d/wright/fulltext.htm 5]
-We did not consider any Amino Acids near the N or C terminus
-We did not consider any residues near ß-sheets or ∂-helices close to the active site because hydrogen bonding plays an active role in substrate stabilization and the polarity of our hix site could disrupt the secondary structure and therefore the hydrogen bonding ability of KNTase)
The yellow bands at the top and bottom of the molecule denotes hix site insertion
We decided to insert our hix sites at the 125 AA of each monomer due to their distance from each other, active site secondary structure, N or C terminus, and lack of any previous mutational analysis proving its function as integral.
Chloramphenicol Acetyltransferase
Cre Recombinase
[http://www3.interscience.wiley.com/cgi-bin/abstract/104558885/ABSTRACT?CRETRY=1&SRETRY=0].