|
Tri-stable Toggle Switch
The purpose of the Tri-stable Toggle Switch is to produce three distinct, continuous, and stable outputs in response to three distinct inputs. These three inputs are three separate chemicals which will each induce one state of the switch. 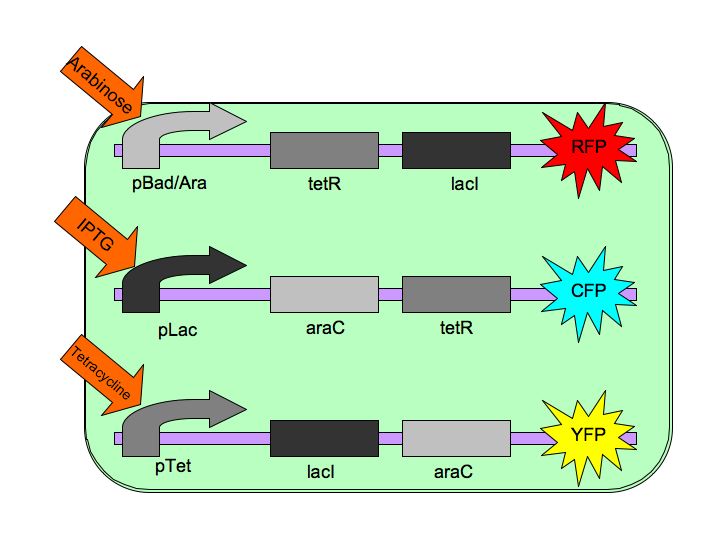 The Tri-stable Toggle Switch Architecture
Each of the three repressors are inactivated by one of three chemicals, the three inducer chemicals mentioned earlier. These three([http://en.wikipedia.org/wiki/Arabinose arabinose], [http://en.wikipedia.org/wiki/IPTG IPTG] (Isopropyl β-D-1-thiogalactopyranoside) and [http://en.wikipedia.org/wiki/Tetracycline Tetracycline], respectively), cause conformational changes in their respective repressor proteins which keeps them from binding to DNA in an inhibitory manner which leads to gene expression. For example, in the presence of arabinose, AraC cannot repress pBAD so LacI and TetR are produced which in turn repress pTet and pLac and the pAraC/BAD construct is turned on.
AraC/BAD
The gene AraC, one of several genes (AraA, AraB, AraD, etc) originally for the metabolism of arabinose.[http://www.mun.ca/biochem/courses/3107/Topics/Ara_operon.html]
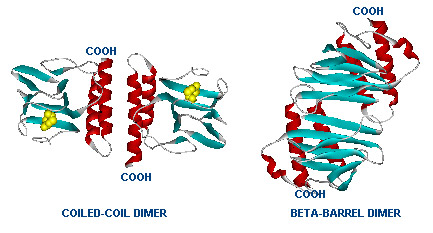 Dimer structure with arabinose on the left (yellow) 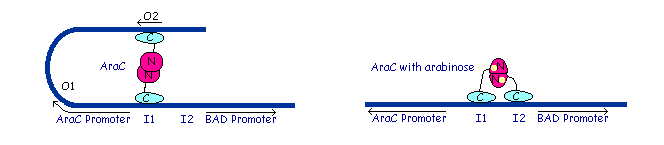 The left image shows the araC dimer repressing transcription, while the right conformation enables transcription
LacI
In nature, LacI represses pLac which promotes the LacYZA genes that metabolize lactose. Thus LacI represses pLac except in the presence of lactose (or lactose mimics, eg IPTG). 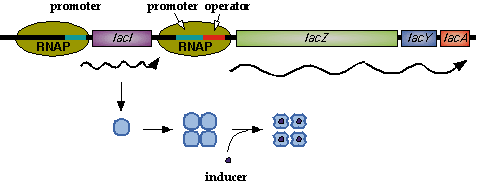 Image[http://www.mun.ca/biochem/courses/3107/Topics/Lac_genetics.html]. LacI forms a tetramer and represses pLac. However, an inducer, such as IPTG, causes a conformation change that removes LacI from the operator site.
TetR
TetR represses the constitutive promoter pTet. In the presence of tetracycline, an antibiotic, a conformational change in TetR inhibits the protein from binding to the operator region. In nature, pTet promotes TetR and TetA. The latter of which acts to pump tetracycline out of the cell, thus the pump is only activated in the presence of Tetracycline.[http://en.wikipedia.org/wiki/Tetracycline_controlled_transcriptional_activation]
The TetR, as it turns out is a very tight repressor and a range of 0 to 1 ug/ml has been shown to cause a 5 order of magnitude change in luciferase production.[http://www.ncbi.nlm.nih.gov/sites/entrez?db=pubmed&cmd=Retrieve&dopt=AbstractPlus&list_uids=1319065&query_hl=1&itool=pubmed_docsum]
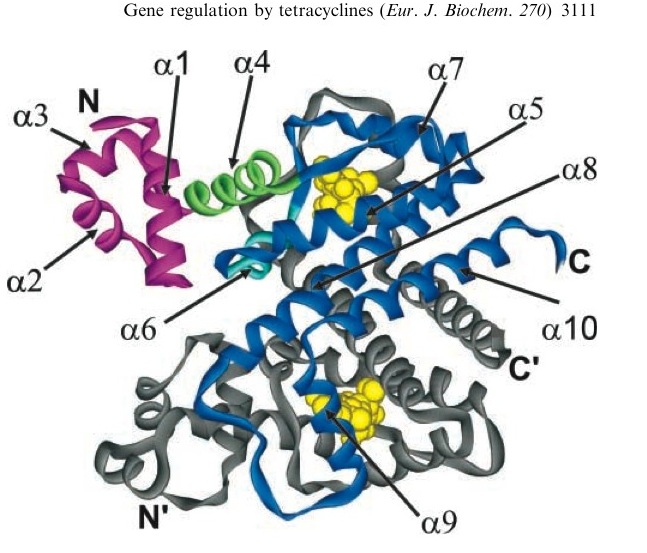 A tetracycline molecule binds to each of the two TetR monomers to form a dimer
Modeling
The first draft of the code started last year:
[http://parts2.mit.edu/wiki/index.php/Table_of_preliminary_model_constants Initial Table of Constants]
[http://parts2.mit.edu/wiki/index.php/Derivation_of_the_Model_Equations Derivation of Model Equations]
Media:tristable2006.txt This code proved to complicated to work with and a more simplified version was developed.
A simpler model based on the bistable paper was developed that takes the combined, relative transcription/translation rates into account Media:tristable2007.txt.
Parameters in the Model
Repressor Production Rate
Repressor production rate is determined by two factors:
- Promoter Strength (Transcription)
- Ribosome Binding Strength (Translation)
In the model, the total repressor production rate = alpha
The promoter strength cannot be easily changed because this would require mutations in the promoter or a different promoter/repressor combo. However, RBSs have been well characterized and the alpha parameter can be modulated by inserting different strength RBSs as determined by the model and testing.
Repressor Strength
The strength of the repressors is determined by:
- The repressor concentration, [Repressor].
- The cooperativity of repression, Beta.
In the Model, the repressor strength = [Repressor]^Beta
The Beta value is characteristic of each repressor and represents a constant that cannot be changed.
|

