Davidson Missouri W/Solving the HPP in vivo
From 2007.igem.org
(→Detecting a Solution) |
|||
| (46 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | <center>[[Davidson Missouri W| <span style="color:red">Home</span>]] | [[Davidson Missouri W/Background Information| <span style="color:red">Background Information</span>]] | [[Davidson Missouri W/Solving the HPP in vivo| <span style="color:black">Current Project: Solving the Hamiltonian Path Problem ''in vivo''</span>]] | [[Davidson Missouri W/Mathematical Modeling| <span style="color:red">Mathematical Modeling</span>]] | [[Davidson Missouri W/Gene splitting| <span style="color:red"> Gene Splitting </span>]] | [[Davidson Missouri W/ | + | <center>[[Davidson Missouri W| <span style="color:red">Home</span>]] | [[Davidson Missouri W/Background Information| <span style="color:red">Background Information</span>]] | [[Davidson Missouri W/Solving the HPP in vivo| <span style="color:black">Current Project: Solving the Hamiltonian Path Problem ''in vivo''</span>]] | [[Davidson Missouri W/Mathematical Modeling| <span style="color:red">Mathematical Modeling</span>]] | [[Davidson Missouri W/Gene splitting| <span style="color:red"> Gene Splitting </span>]] | [[Davidson Missouri W/Results| <span style="color:red">Results</span>]] | [[Davidson Missouri W/Traveling Salesperson Problem| <span style="color:red">Traveling Salesperson Problem</span> ]] | [[Davidson Missouri W/Software|<span style="color:red">Software</span>]] | [[Davidson Missouri W/Resources and Citations|<span style="color:red">Resources and Citations</span>]]</center> |
<hr> | <hr> | ||
| - | + | <br> | |
| - | Using the Hin/'' | + | Using the Hin/''hixC'' flipping mechanism, we are developing a bacterial computer which solves a specific mathematical problem, the ''Hamiltonian Path'' problem. |
=The Hamiltonian Path Problem= | =The Hamiltonian Path Problem= | ||
| - | A Hamiltonian Path is a trip through a graph which visits each node exactly once. A graph may have multiple Hamiltonian Paths, only one, or even none. | + | A Hamiltonian Path is a trip through a directed graph which visits each node exactly once. A graph may have multiple Hamiltonian Paths, only one, or even none. The problem asks the following: given a graph with a designated starting point and an endpoint, does it contain a Hamiltonian path? |
We solve our problem by transforming ''E. coli'' cells with specially engineered plasmids. | We solve our problem by transforming ''E. coli'' cells with specially engineered plasmids. | ||
==Designing a Plasmid== | ==Designing a Plasmid== | ||
| - | Our plasmid consists of reporter genes and '' | + | Our plasmid consists of reporter genes and ''hixC'' sites. ''hixC'' sites are placed within the coding regions of our reporter genes. The reporter genes are joined in such a way as to represent a graph. Each reporter gene represents a node, and the connection of two different reporter gene halves together without any ''hixC'' sites in between represents an edge. |
| - | [[Image: | + | Such a plasmid design allows us to flip the edges of the graph into all possible permutations. Each permutation represents a potential path through the graph; permutations with all the proper phenotypes can be selected for. This system therefore allows for the detection of computation, as solutions to the Hamiltonian Path problem will manifest themselves via the reporter genes. |
| + | |||
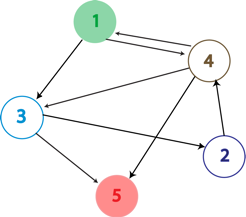
| + | [[Image:Hppgraph.png|frame|left|An arbitrary graph with 5 nodes and 8 edges. We want to determine if a Hamiltonian Path exists in the graph, starting at node 1 (green) and ending at node 5 (red).]] [[Image:Hppplasmid.png|thumb|600px|none|A possible starting construct to find a Hamiltonian Path in the graph to the left. Each edge in the graph is included on the plasmid as a flippable unit that is flanked by ''hixC'' sites. The representation of an edge on the plasmid consists of the back half of the gene you are leaving and the front half of the gene you are going towards. The front half of the starting gene (gene 1) is not flippable and is placed at the beginning of the construct to ensure that it is the first gene in the path. The ending node in the graph (gene 5) would be a transciptional terminator to ensure that it is the final gene in the path.]] | ||
| + | <br> | ||
| + | |||
| + | [[Image:Hppplasmidsolved.png|thumb|600px|none|The Hamiltonian Path solution (post Hin-mediated recombination). Once the starting construct (above) is exposed to Hin Recombinase, flipping of DNA segments that are flanked by ''hixC'' sites will occur. We would select for a plasmid that has solved the problem based on phenotypic expression of all reporter genes.]] | ||
| + | <br> | ||
==Developing Nodes== | ==Developing Nodes== | ||
| - | We represent the graph's nodes with reporter genes. In order to allow for flipping, we must insert '' | + | We represent the graph's nodes with reporter genes. In order to allow for flipping, we must insert ''hixC'' sites within the coding regions of our reporter genes. We call this process [[Davidson_Missouri_W/Gene splitting|''gene splitting'']]. The central issue with gene splitting is that an additional 13 amino acids are added within the resulting protein. More often than not, we would expect these amino acids to prevent the protein from functioning. If our reporter gene tolerates a ''hixC'' insertion, and still functions as a reporter properly, then we can use it as a node on our graph. |
| + | |||
| + | ==Initiating Flipping== | ||
| + | After a plasmid has been constructed with the the reporter genes in a scrambled order, flipping can be initiated by co-transformation with a plasmid containing the Hin protein. The initiation of gene flipping commences computation. Eventually, some gene portions may flip into orientations which are solutions to the Hamiltonian path problem. | ||
| + | |||
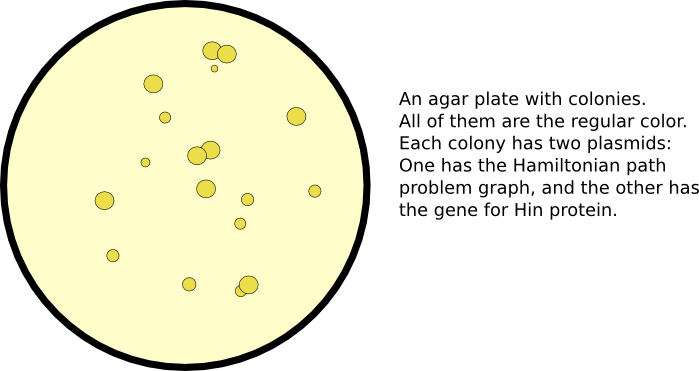
| + | [[Image:Unflipped_agar_plate.png|thumb|600px|none|]] | ||
| + | |||
| + | ==Detecting a Solution== | ||
| + | The primary method for seeing if there is a solution is by screening for all correct '''phenotypes'''. For example, if some cells do not have antibiotic resistance, or if they do not exhibit a particular color, then we know they have not computed a solution to the problem. Our computer constructs are all designed to use the T7 RNA polymerase promoter. This allows us to initiate transcription as a separate event from flipping upon co-transformation with a separate vector containing the gene for the T7 RNA polymerase. Another advantage of using this promoter and polymerase is its supposed greater processivity. Larger graphs require the transcription of more genes; typically, most promoters only promote through a few. Although we have not tested more than two genes with this promoter, we expect that this system permits greater transcriptional power. | ||
| + | |||
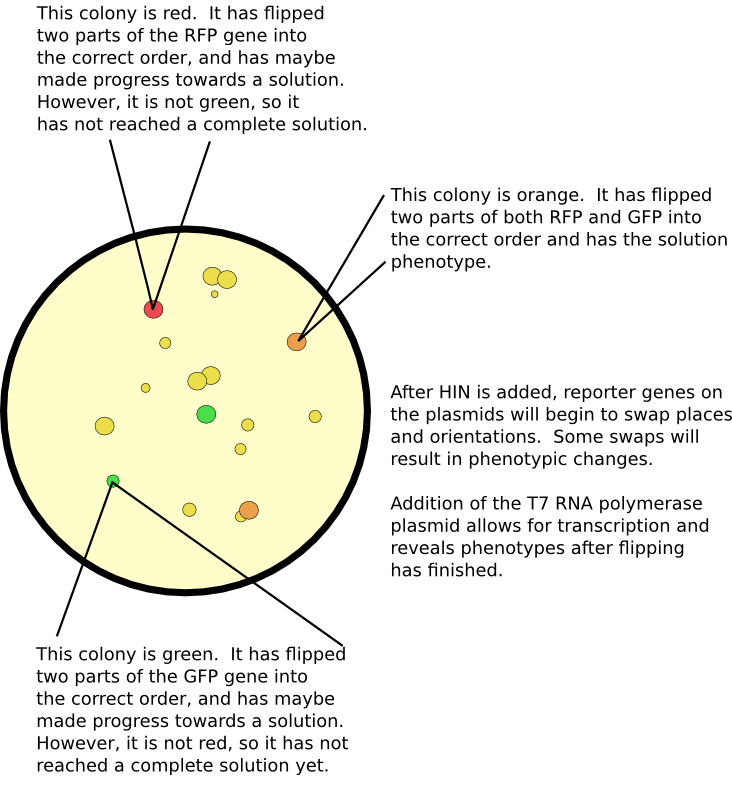
| + | [[Image:Flipped_explain.png|thumb|600px|none|In this example, the two reporter genes are GFP and RFP. Cells with a solution will have both green and red fluorescence (which may appear orange). Cells without a complete solution will not express both proteins.]] | ||
| + | |||
| + | Although phenotypic screening is our primary means of verifying for solutions, there exists the possibility of a '''false-positive''' solution. In this case, all genes have their second portion immediately following their first. Yet the resulting path will be illogical: it will contain a "teleportation," which means that, in order to travel the path, it would be necessary to move between two nodes without following an edge. Go [[Davidson_Missouri_W/Mathematical_Modeling#True_Positives_and_False_Positives | here]] for more information on false positives. | ||
| + | |||
| + | The potential for false-positives thus requires another way of verifying solutions. The length of a solution, in terms of base-pairs, is always known. It is the sum of the lengths of all split genes. Furthermore, the first and last nodes are constant, as the first one is not flippable and the last node is the transcriptional terminator. These facts mean that '''PCR''' can be used to verify solutions. Primers can be designed from the first and last node, and running PCR products on a gel will give the lengths of the amplified DNA. If any segment has the correct length then we know there is a solution. | ||
| + | |||
| + | <html> | ||
| + | <head> | ||
| + | <link rel="stylesheet" href="https://2007.igem.org/wiki/index.php?title=User:Wideloache/igem2007.css&action=raw&ctype=text/css" type="text/css" /> | ||
| + | </style> | ||
| + | </head> | ||
| + | <body> | ||
| + | |||
| + | <div id="roll"> | ||
| + | <img src="https://static.igem.org/mediawiki/2007/3/30/Agarose_gel_b.png" alt="Agarose Gel with PCR Products"> | ||
| + | <ul> | ||
| + | <li id="true"><a href=""></a></li> | ||
| + | <li id="false"><a href=""></a></li> | ||
| + | <li id="negative"><a href=""></a></li> | ||
| + | </ul> | ||
| + | </div> | ||
| + | |||
| + | </body> | ||
| + | </html> | ||
| + | |||
| + | <br> | ||
| + | <hr> | ||
| + | <center> | ||
| + | <[[Davidson Missouri W/Background Information | Previous Section]] | [[Davidson Missouri W/Mathematical Modeling | Next Section>]] | ||
| + | </center> | ||
Latest revision as of 18:22, 23 October 2007
Using the Hin/hixC flipping mechanism, we are developing a bacterial computer which solves a specific mathematical problem, the Hamiltonian Path problem.
Contents |
The Hamiltonian Path Problem
A Hamiltonian Path is a trip through a directed graph which visits each node exactly once. A graph may have multiple Hamiltonian Paths, only one, or even none. The problem asks the following: given a graph with a designated starting point and an endpoint, does it contain a Hamiltonian path?
We solve our problem by transforming E. coli cells with specially engineered plasmids.
Designing a Plasmid
Our plasmid consists of reporter genes and hixC sites. hixC sites are placed within the coding regions of our reporter genes. The reporter genes are joined in such a way as to represent a graph. Each reporter gene represents a node, and the connection of two different reporter gene halves together without any hixC sites in between represents an edge.
Such a plasmid design allows us to flip the edges of the graph into all possible permutations. Each permutation represents a potential path through the graph; permutations with all the proper phenotypes can be selected for. This system therefore allows for the detection of computation, as solutions to the Hamiltonian Path problem will manifest themselves via the reporter genes.
Developing Nodes
We represent the graph's nodes with reporter genes. In order to allow for flipping, we must insert hixC sites within the coding regions of our reporter genes. We call this process gene splitting. The central issue with gene splitting is that an additional 13 amino acids are added within the resulting protein. More often than not, we would expect these amino acids to prevent the protein from functioning. If our reporter gene tolerates a hixC insertion, and still functions as a reporter properly, then we can use it as a node on our graph.
Initiating Flipping
After a plasmid has been constructed with the the reporter genes in a scrambled order, flipping can be initiated by co-transformation with a plasmid containing the Hin protein. The initiation of gene flipping commences computation. Eventually, some gene portions may flip into orientations which are solutions to the Hamiltonian path problem.
Detecting a Solution
The primary method for seeing if there is a solution is by screening for all correct phenotypes. For example, if some cells do not have antibiotic resistance, or if they do not exhibit a particular color, then we know they have not computed a solution to the problem. Our computer constructs are all designed to use the T7 RNA polymerase promoter. This allows us to initiate transcription as a separate event from flipping upon co-transformation with a separate vector containing the gene for the T7 RNA polymerase. Another advantage of using this promoter and polymerase is its supposed greater processivity. Larger graphs require the transcription of more genes; typically, most promoters only promote through a few. Although we have not tested more than two genes with this promoter, we expect that this system permits greater transcriptional power.
Although phenotypic screening is our primary means of verifying for solutions, there exists the possibility of a false-positive solution. In this case, all genes have their second portion immediately following their first. Yet the resulting path will be illogical: it will contain a "teleportation," which means that, in order to travel the path, it would be necessary to move between two nodes without following an edge. Go here for more information on false positives.
The potential for false-positives thus requires another way of verifying solutions. The length of a solution, in terms of base-pairs, is always known. It is the sum of the lengths of all split genes. Furthermore, the first and last nodes are constant, as the first one is not flippable and the last node is the transcriptional terminator. These facts mean that PCR can be used to verify solutions. Primers can be designed from the first and last node, and running PCR products on a gel will give the lengths of the amplified DNA. If any segment has the correct length then we know there is a solution.