Wisconsin/Project
From 2007.igem.org
Sean RM177 (Talk | contribs) (→Experiment) |
|||
| (4 intermediate revisions not shown) | |||
| Line 6: | Line 6: | ||
Artificial transcription factor (ATF) has two domains: DNA-binding domain and regulatory domain. The DNA-binding domain provides sequence specific targeting and the regulatory domain can up- or down-regulate gene expression. ATF design is modular, meaning one can mix and match different DNA-binding motifs with various transcription factors. | Artificial transcription factor (ATF) has two domains: DNA-binding domain and regulatory domain. The DNA-binding domain provides sequence specific targeting and the regulatory domain can up- or down-regulate gene expression. ATF design is modular, meaning one can mix and match different DNA-binding motifs with various transcription factors. | ||
| - | We decided to use zinc fingers for DNA-binding. A zinc finger is made of about 30 amino acids and is stabilized by a single zinc ion to form a certain protein structure (eg. beta-beta-alpha). It can interact with a specific 3 base-pair motif in DNA (Segal & Barbas, 2001). Regulation of gene activity is achieved by using zinc finger to recognize a specific DNA sequences | + | We decided to use zinc fingers for the DNA-binding domain. A zinc finger is made of about 30 amino acids and is stabilized by a single zinc ion to form a certain protein structure (eg. beta-beta-alpha). It can interact with a specific 3 base-pair motif in DNA (Segal & Barbas, 2001). Regulation of gene activity is achieved by using zinc finger to recognize and bind to a specific DNA sequences, and then recruite other molecular machinery that will activate or repress the gene. Our ATF project has focused on characterizing DNA binding specificity and cooperativity effects. |
===Design=== | ===Design=== | ||
| - | Our ATF design has 3 zinc fingers | + | Our ATF design has 3 zinc fingers, each can recognize 3bp of DNA sequence. So it should bind to a unique 9bp sequence. Following the zinc fingers are NLS and VP64. NLS is for localizing ATF to the nucleus and VP64 is a strong transcription activator. In order to test DNA-binding specificity, we also performed site-directed mutagenesis to change the zinc finger 1's recognition sequence from GAA to GGG (as shown below). This way we can test how well similar zinc fingers can discriminate DNA sequences. |
[[image:zif.jpg]] | [[image:zif.jpg]] | ||
===Cognate Site Identity (CSI) Microarray=== | ===Cognate Site Identity (CSI) Microarray=== | ||
| - | We are using CSI microarrays to quantify how well zinc fingers bind to DNA sequences. The CSI array contains every permutation of 9bp DNA sequence. This allows us to test the entire sequence space in an unbiased manner. | + | We are using CSI microarrays to quantify how well zinc fingers bind to DNA sequences. The CSI array we designed contains every permutation of 9bp DNA sequence. This allows us to test the entire sequence space in an unbiased manner. Below is a preliminary CSI array scan of purified ATF protein on an 8mer CSI array. |
| + | |||
| + | [[Image:csiarray.jpg]] | ||
==BCL-2 and Cell Fate Regulation== | ==BCL-2 and Cell Fate Regulation== | ||
===Background=== | ===Background=== | ||
| - | Doxrubicin is a widely used drug to treat cancer. However one | + | Doxrubicin is a widely used drug to treat cancer. However one of its main limitation is cardiotoxicity. Accumulated dose of doxrubicin can cause oxidative stress to cardiomyocytes and trigger apoptosis. Our goal is to use ATF to up-regulate BCL-2 (a pro-survival protein) and prevent cells from dying under mild stress. |
===Design=== | ===Design=== | ||
| - | + | We designed a zinc finger ATF that will bind to BCL-2 promoter region and up-regulate transcription. The mammalian expression vector we used is tetracycline controlled. Adding tetracycline will induce ATF expression. | |
[[Image:tetsys.jpg]] | [[Image:tetsys.jpg]] | ||
===Experiment=== | ===Experiment=== | ||
| - | We tested our construct for function in | + | We tested our construct for function in NIH 3T3 (mouse fibroblast) cells. The first part of our experiment is to check for ATF and BCL-2 expression. We did this by looking at western blots. We successfully transfected our ATF into fibroblast cells but we were unable to detect any ATF proteins. We tried using different concentration of tetracycline but that did not improve our results. Since ATF was not expressed, the BCL-2 levels remained constant under all conditions. This is expected since BCL-2 is constitutively expressed under normal conditions. Unfortunately we were not able to test how BCL-2 levels affect cell fate regulation. |
Latest revision as of 00:52, 27 October 2007
Contents |
Project Overview
Reprogramming gene expression is one of the main goals of synthetic biology. In the past, most groups have focused on cis-regulatory elements such as promoter to control transcription. We decided to explore artificial transcription factor (ATF), a trans-regulatory element, for controlling genes. Our project consists of characterizing ATF and testing ATF in mammalian cells.
Artificial Transcription Factor
Background
Artificial transcription factor (ATF) has two domains: DNA-binding domain and regulatory domain. The DNA-binding domain provides sequence specific targeting and the regulatory domain can up- or down-regulate gene expression. ATF design is modular, meaning one can mix and match different DNA-binding motifs with various transcription factors.
We decided to use zinc fingers for the DNA-binding domain. A zinc finger is made of about 30 amino acids and is stabilized by a single zinc ion to form a certain protein structure (eg. beta-beta-alpha). It can interact with a specific 3 base-pair motif in DNA (Segal & Barbas, 2001). Regulation of gene activity is achieved by using zinc finger to recognize and bind to a specific DNA sequences, and then recruite other molecular machinery that will activate or repress the gene. Our ATF project has focused on characterizing DNA binding specificity and cooperativity effects.
Design
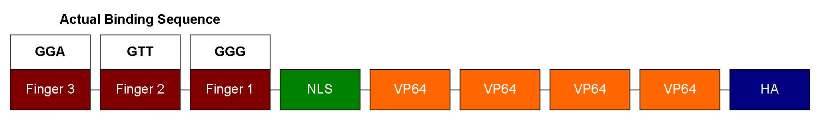
Our ATF design has 3 zinc fingers, each can recognize 3bp of DNA sequence. So it should bind to a unique 9bp sequence. Following the zinc fingers are NLS and VP64. NLS is for localizing ATF to the nucleus and VP64 is a strong transcription activator. In order to test DNA-binding specificity, we also performed site-directed mutagenesis to change the zinc finger 1's recognition sequence from GAA to GGG (as shown below). This way we can test how well similar zinc fingers can discriminate DNA sequences.
Cognate Site Identity (CSI) Microarray
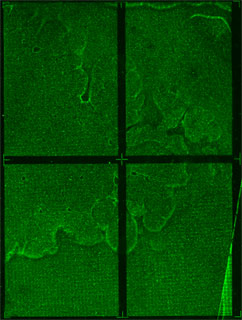
We are using CSI microarrays to quantify how well zinc fingers bind to DNA sequences. The CSI array we designed contains every permutation of 9bp DNA sequence. This allows us to test the entire sequence space in an unbiased manner. Below is a preliminary CSI array scan of purified ATF protein on an 8mer CSI array.
BCL-2 and Cell Fate Regulation
Background
Doxrubicin is a widely used drug to treat cancer. However one of its main limitation is cardiotoxicity. Accumulated dose of doxrubicin can cause oxidative stress to cardiomyocytes and trigger apoptosis. Our goal is to use ATF to up-regulate BCL-2 (a pro-survival protein) and prevent cells from dying under mild stress.
Design
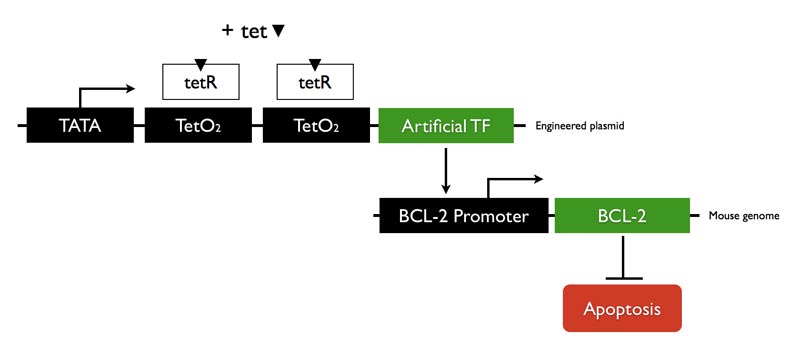
We designed a zinc finger ATF that will bind to BCL-2 promoter region and up-regulate transcription. The mammalian expression vector we used is tetracycline controlled. Adding tetracycline will induce ATF expression.
Experiment
We tested our construct for function in NIH 3T3 (mouse fibroblast) cells. The first part of our experiment is to check for ATF and BCL-2 expression. We did this by looking at western blots. We successfully transfected our ATF into fibroblast cells but we were unable to detect any ATF proteins. We tried using different concentration of tetracycline but that did not improve our results. Since ATF was not expressed, the BCL-2 levels remained constant under all conditions. This is expected since BCL-2 is constitutively expressed under normal conditions. Unfortunately we were not able to test how BCL-2 levels affect cell fate regulation.