Chiba/Engeneering Flagella
From 2007.igem.org
(→Results&Discussion) |
(→Our Aim) |
||
| (57 intermediate revisions not shown) | |||
| Line 2: | Line 2: | ||
[[Image:chiba_logo.png|center]] | [[Image:chiba_logo.png|center]] | ||
__NOTOC__ | __NOTOC__ | ||
| - | {| style="border:0;width:100%;" cellpadding="20px" cellspacing="0" | + | {| style="border:0;width:100%;font-family:'Trebuchet MS'" cellpadding="20px" cellspacing="0" |
| align="center" | | | align="center" | | ||
| - | [[Chiba|Introduction]] | [[Chiba/Project_Design|Project Design]] ( [[Chiba/Engeneering_Flagella|1.Affinity Tag]] | [[Chiba/Communication|2.Communication Module]] | [[Chiba/Quorum_Sensing|3.Size Control]] ) | [[Chiba/Making Marimo|Making Marimos]] | [[Chiba/Goal|Our Goal]]<br> | + | [[Chiba|Home]]<br> |
| + | <span style="font-size:120%;font-weight:bold;">[[Chiba/Introduction|Introduction]] | [[Chiba/Project_Design|Project Design]] ( [[Chiba/Engeneering_Flagella|1.Affinity Tag]] | [[Chiba/Communication|2.Communication Module]] | [[Chiba/Quorum_Sensing|3.Size Control]] ) | [[Chiba/Making Marimo|Making Marimos]] | [[Chiba/Goal|Our Goal]]</span><br> | ||
[[Chiba/Acknowledgements|Acknowledgements]] | [[Chiba/Team_Members|Team Members]] | [http://chem.tf.chiba-u.jp/igem/ iGEM Chiba Website] | [[Chiba/Members_Only|メンバ連絡簿]] | [[Chiba/Acknowledgements|Acknowledgements]] | [[Chiba/Team_Members|Team Members]] | [http://chem.tf.chiba-u.jp/igem/ iGEM Chiba Website] | [[Chiba/Members_Only|メンバ連絡簿]] | ||
|} | |} | ||
| - | + | ==Affinity Tags== | |
| - | == | + | |
===Our Aim=== | ===Our Aim=== | ||
| - | [[Image:Chiba_stickbacteria.png|frame|''' | + | [[Image:Chiba_stickbacteria.png|frame|'''Fig. 9''' Affinity tags.]] |
| - | To make | + | To make affinity tags on ''E.coli'', we focused on their flagella that are located outside the cells. We used the following mechanisms: |
*Display sticky peptides in flagellar filament. | *Display sticky peptides in flagellar filament. | ||
*His-tag. The imidazole group in histidines make a complex with metal ions. | *His-tag. The imidazole group in histidines make a complex with metal ions. | ||
| Line 24: | Line 24: | ||
The filament of ''E.Coli'' is a rigid, helical, and cylindrical structure which is 10-15μm long and 23nm thick in diameter. | The filament of ''E.Coli'' is a rigid, helical, and cylindrical structure which is 10-15μm long and 23nm thick in diameter. | ||
It is built from ~20000 subunits of a ~55kDa single protein, FliC. | It is built from ~20000 subunits of a ~55kDa single protein, FliC. | ||
| - | FliC has three domains, D1,D2,D3; although D1 and D2 are needed for the formation of the functional flagellar filament, D3 domain which sticks outside of the fillament are not essential[ | + | FliC has three domains, D1,D2,D3; although D1 and D2 are needed for the formation of the functional flagellar filament, D3 domain which sticks outside of the fillament are not essential[3]. |
===="Variable" FliC D3 domain==== | ===="Variable" FliC D3 domain==== | ||
| - | It is reported that the proteins up to 49.4kDa could be displayed on the cell surface of ''E.Coli'' using flagellin fusion protein.[ | + | It is reported that the proteins up to 49.4kDa could be displayed on the cell surface of ''E.Coli'' using flagellin fusion protein.[4] |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
===About Histidine Tag=== | ===About Histidine Tag=== | ||
| Line 40: | Line 36: | ||
*We inserted the short peptide with six histidine (“His-Tag”) into the fliC D3 domain. | *We inserted the short peptide with six histidine (“His-Tag”) into the fliC D3 domain. | ||
| - | [[Image:Chiba_flichisgene.png]]<br> | + | [[Image:Chiba_flichisgene.png|frame|'''Fig.10''' flic his gene]]<br> |
*[[Image:Chiba_check.png]] Sequence Confirmed<br> | *[[Image:Chiba_check.png]] Sequence Confirmed<br> | ||
| Line 51: | Line 47: | ||
====Samples==== | ====Samples==== | ||
| - | * | + | *⊿fliC strain(JW1908 in KEIO collections [5]) transformed with |
**pUC19-fliC-his | **pUC19-fliC-his | ||
**no plasmid | **no plasmid | ||
| - | * | + | *⊿fliC,⊿motB strain(GI826)transformed with |
**pUC19-fliC-his | **pUC19-fliC-his | ||
**no plasmid | **no plasmid | ||
| + | <br> | ||
====Testing Procedure==== | ====Testing Procedure==== | ||
| - | #pUC19-FliC-His was transformed to JW1908(''fliC'') and GI826(''fliC'' ''motB''). | + | #pUC19-FliC-His was transformed to JW1908(''fliC''), JW0747(''MotB''), and GI826(''fliC'' ''motB''). |
#Grown to stationary phase | #Grown to stationary phase | ||
#Culture suspended with Dynabeads (Metal-IDA), allowing to the affinity adsorption | #Culture suspended with Dynabeads (Metal-IDA), allowing to the affinity adsorption | ||
| Line 67: | Line 64: | ||
====Results&Discussion==== | ====Results&Discussion==== | ||
| - | 1.Stickiness check | + | '''1.''Stickiness'' check using FliC strain''' |
| - | [[Image:Chiba Beads-Adsorption result3.png|frame|left| | + | [[Image:Chiba Beads-Adsorption result3.png|frame|left|'''Fig. 11''' Binding test using Strain JW1908(''⊿fliC'',).]]<br clear="all"> |
| - | [[Image:|frame|left| | + | *Cell without His-FliC bound better to the Beads? No way! |
| - | * | + | *We thought the problem might be the super-fast revolution of flagella itself. We decided to try MotB strain.<br><br><br> |
| - | Co<sup>2+</sup> | + | |
| - | + | '''2.''Stickiness'' check using MotB strain''' | |
| - | * | + | [[Image:Chiba Beads-Adsorption result2.png|frame|left|'''fig. 12''' Binding test using Strain JW0747(''⊿motB'').]]<br clear="all"> |
| - | * | + | *Mot B deletion provides cell with the flagella completely assembled but not rotating. <br> |
| + | *'''This time everything worked out!''' Only in the presence of Co<sup>2+</sup>Bacteria with His-FliC sticked to the Co-IDA beads very well. | ||
| + | *In this strain, FliC-His is assembled with wildtype FliC coded in genomic DNA. Nevertheless, the binding efficiency was at the same level (not shown). it seems that His-Tag displayed on the flagella is enough to do its work. | ||
| + | *On the other hand, the deletion of MotB turned out to be vital for sticking the tagged flagella together. | ||
| + | *In the presence of FliC-His, cobalt ion adsorb bacteria stronger than nickel ion, this was more or less the expected result. | ||
| + | |||
| + | |||
<!-- | <!-- | ||
| Line 83: | Line 86: | ||
--> | --> | ||
| - | + | ==References== | |
| - | + | 3. Kuwajima, G. ''et al''.: ''J. Bacteriol.'', '''170''', 3305-3309 (1988)<br> | |
| - | <br> | + | 4. Ezaki, S. ''et. al''.: ''J. Ferment. Bioeng.'', '''86''', 500-503 (1998)<br> |
| - | + | 5. Baba, T. ''et. al''.: ''Mol. Systems. Biol.,'' '''21''', 1-10 (2006)<br> | |
| - | + | ||
| - | + | ||
Latest revision as of 05:26, 27 October 2007
|
Home |
Affinity Tags
Our Aim
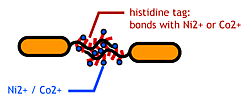
To make affinity tags on E.coli, we focused on their flagella that are located outside the cells. We used the following mechanisms:
- Display sticky peptides in flagellar filament.
- His-tag. The imidazole group in histidines make a complex with metal ions.
We combined these two and made a His-tagged flagella in the hope to stick them together via metal ions.
[http://www.npn.jst.go.jp/index.html About flagella]
E.Coli have 5-10 flagella. The flagella is used for swimming and for chemotaxis; the bacteria run when they find attractant, tumble when there is a repellent.
E.coli flagella consist of three parts: a basal body, a hook, and a filament. The filament of E.Coli is a rigid, helical, and cylindrical structure which is 10-15μm long and 23nm thick in diameter. It is built from ~20000 subunits of a ~55kDa single protein, FliC. FliC has three domains, D1,D2,D3; although D1 and D2 are needed for the formation of the functional flagellar filament, D3 domain which sticks outside of the fillament are not essential[3].
"Variable" FliC D3 domain
It is reported that the proteins up to 49.4kDa could be displayed on the cell surface of E.Coli using flagellin fusion protein.[4]
About Histidine Tag
See [http://en.wikipedia.org/wiki/His-tag wikipedia article].
Experiments
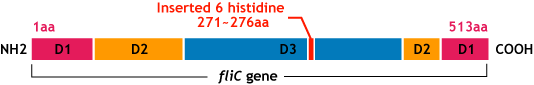
Making FliC-his gene
- We inserted the short peptide with six histidine (“His-Tag”) into the fliC D3 domain.
Checking the "Stickiness": Beads Adsorption
Purpose
Confirm that the his-tags are displaied on the flagella and are capable of binding to Co2+- or Ni2+- surface.
Samples
- ⊿fliC strain(JW1908 in KEIO collections [5]) transformed with
- pUC19-fliC-his
- no plasmid
- ⊿fliC,⊿motB strain(GI826)transformed with
- pUC19-fliC-his
- no plasmid
Testing Procedure
- pUC19-FliC-His was transformed to JW1908(fliC), JW0747(MotB), and GI826(fliC motB).
- Grown to stationary phase
- Culture suspended with Dynabeads (Metal-IDA), allowing to the affinity adsorption
- Beads washed with a phosphate buffer (x4)
- E" coli" detached from beads by adding imidazole then spreaded on agar plates.
- The number of the colonies on resultant plates.
Results&Discussion
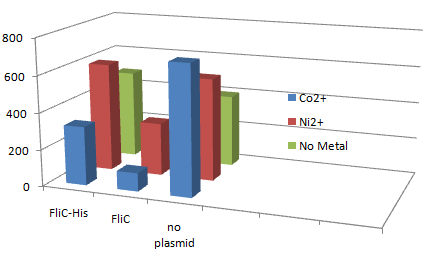
1.Stickiness check using FliC strain
- Cell without His-FliC bound better to the Beads? No way!
- We thought the problem might be the super-fast revolution of flagella itself. We decided to try MotB strain.
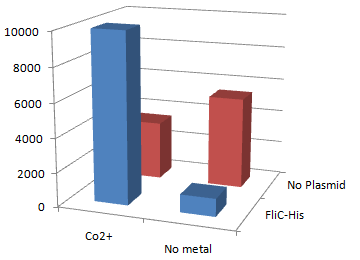
2.Stickiness check using MotB strain
- Mot B deletion provides cell with the flagella completely assembled but not rotating.
- This time everything worked out! Only in the presence of Co2+Bacteria with His-FliC sticked to the Co-IDA beads very well.
- In this strain, FliC-His is assembled with wildtype FliC coded in genomic DNA. Nevertheless, the binding efficiency was at the same level (not shown). it seems that His-Tag displayed on the flagella is enough to do its work.
- On the other hand, the deletion of MotB turned out to be vital for sticking the tagged flagella together.
- In the presence of FliC-His, cobalt ion adsorb bacteria stronger than nickel ion, this was more or less the expected result.
References
3. Kuwajima, G. et al.: J. Bacteriol., 170, 3305-3309 (1988)
4. Ezaki, S. et. al.: J. Ferment. Bioeng., 86, 500-503 (1998)
5. Baba, T. et. al.: Mol. Systems. Biol., 21, 1-10 (2006)