Rice
From 2007.igem.org
| Line 40: | Line 40: | ||
='''2007 Team Roster'''= | ='''2007 Team Roster'''= | ||
[[Image:RiceSyntheticTitle.jpg|center]] | [[Image:RiceSyntheticTitle.jpg|center]] | ||
| - | <div | + | <BR><BR> |
| - | [[Image:Rice07Team.jpg|thumb|300px|Top: Thomas, Taylor, David. | + | <div style= "position: absolute; left:500px"> |
| + | [[Image:Rice07Team.jpg|thumb|none|300px|Top: Thomas, Taylor, David. | ||
Bottom: Joff, Peter, Beth, and Alec.]] | Bottom: Joff, Peter, Beth, and Alec.]] | ||
</div> | </div> | ||
| - | <div style=" | + | <div style= "position: absolute; left:200px"> |
* '''Undergraduate Students''' | * '''Undergraduate Students''' | ||
** Tina Chen | ** Tina Chen | ||
| Line 51: | Line 52: | ||
** David Kim | ** David Kim | ||
** Alec Walker | ** Alec Walker | ||
| - | + | <BR> | |
* '''Graduate Advisors''' | * '''Graduate Advisors''' | ||
** Bibhash Mukhopadhyay | ** Bibhash Mukhopadhyay | ||
| Line 57: | Line 58: | ||
** Christie Peebles | ** Christie Peebles | ||
** Jay Raol | ** Jay Raol | ||
| - | + | <BR> | |
* '''Faculty Advisors''' | * '''Faculty Advisors''' | ||
| - | ** Beth Beason | + | ** Beth Beason, PhD. |
| - | ** George Bennett | + | ** George Bennett, PhD. |
| - | ** Steve Cox | + | ** Steve Cox, PhD. |
| - | ** Ken Cox | + | ** Ken Cox, PhD. |
| - | ** Ka-yiu San | + | ** Ka-yiu San, PhD. |
| - | ** Joff Silberg | + | ** Joff Silberg, PhD. |
</div> | </div> | ||
</font> | </font> | ||
Revision as of 17:59, 26 October 2007
Welcome to Rice University's 2007 iGEM wiki page. We have a number of incredible projects this year! Our new project is focused on the design a circuit to modulate phage-bacteria ecological dynamics towards the goal of selecting against antibiotic resistance. The continuation of last year's project involves the construction of a novel bacterial phenotype, quorumtaxis, in which bacteria are programmed to detect and pursue other bacterial species.
Contents |
Project A: Phage Project Overview
Recently it has been demonstrated how differential selection between antibiotic resistance and sensitive bacterial populations is affected under a regime of multiple drug combinations. Normally, if two populations of the same bacterial strain compete, the one which mutated to acquire anti-biotic resistance 'wins' the fight for survival over the ones that remained wild-type and antibiotic sensitive. However, in certain drug combination regimens that involve suppressive interactions, high drug concentrations results in competitive selection against resistance, without perturbing the effectiveness of the other drug in the combination. These results are interpreted as coming from a trade-off regime in the evolution of resistance fitness landscape, where absolute potency of a drug combination is balanced by the relative competitive selection imposed on emerging resistant populations.
Following up on the elucidation of this phenomenon, our iGEM team asks the following question: Can we artificially engineer bacteriophage which would produce a selection against antibiotic resistance within bacterial populations?
Details...
Project B: Quorumtaxis Project Overview
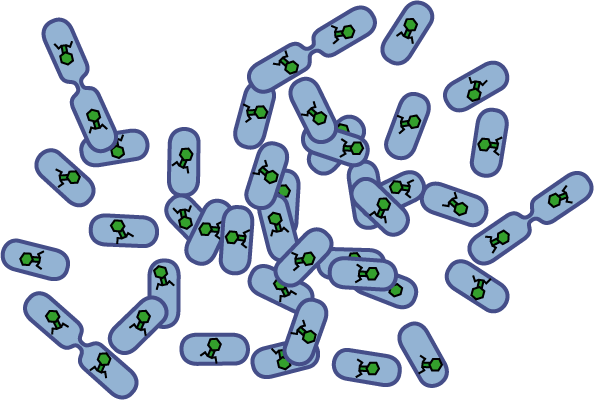
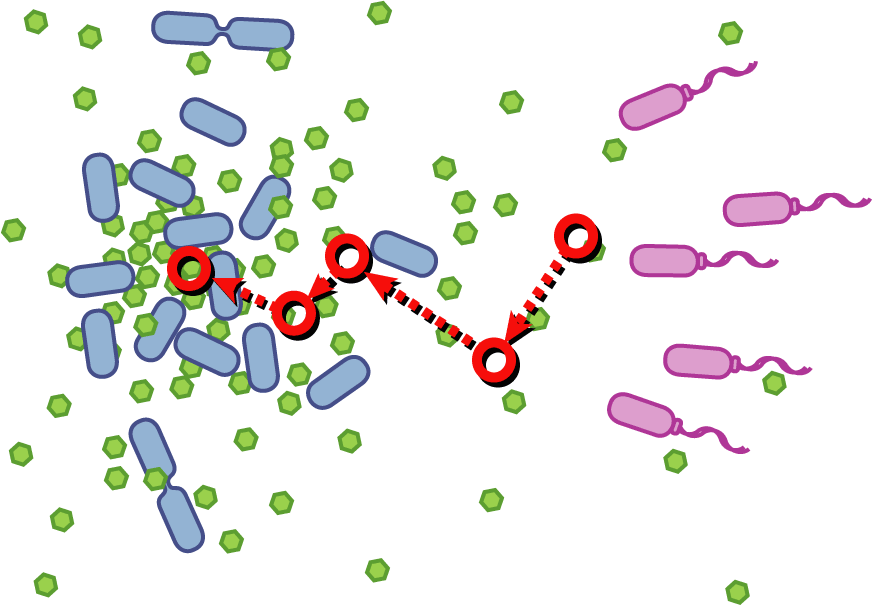
Bacteria have evolved diverse genetic systems to sense their environment as well as respond to their surroundings in an adaptive manner. Of recent intense interest is the discovery that bacteria use pheromones that allow them to sense their own population density, a system known as quorum sensing. Another widely studied system is that of chemotaxis, in which a bacterium is able to sense and adaptively swim towards or away from a chemical agent in the environment. The Rice University iGEM team is attempting to merge these two existing natural systems (quorum sensing and chemotaxis) to produce a novel bacterial phenotype (quorumtaxis) in which the engineered cell will be able to detect and swim towards the quorum pheromone of another ‘target’ cell. This project will demonstrate the ability to produce unique complex behavior in bacteria through the modular integration of existing circuits. In addition, precise control over bacterial movement will greatly increase the complexity of systems synthetic biologists could create. The keystone of the project is the design of a novel chimeric receptor which can sense the target pheromone and then signal flagellar rotation within the cell. This project necessitates a highly interdisciplinary approach: the conceptual design and part construction requires backgrounds in biochemistry and cell biology, protein engineering elements requires experience in biomolecular engineering methods, and computational mathematics will be used to model the quorumtaxis phenotype.
Our iGEM team asks the following question: Can we re-engineer E.coli to recognize and pursue other bacterial species?
Details...
Thanks for visiting our wiki, and be sure to tell us what you think when we see you at the iGEM Jamboree!
2007 Team Roster
- Undergraduate Students
- Tina Chen
- Thomas Segall-Shapiro
- Taylor Stevenson
- David Kim
- Alec Walker
- Graduate Advisors
- Bibhash Mukhopadhyay
- Peter Nguyen
- Christie Peebles
- Jay Raol
- Faculty Advisors
- Beth Beason, PhD.
- George Bennett, PhD.
- Steve Cox, PhD.
- Ken Cox, PhD.
- Ka-yiu San, PhD.
- Joff Silberg, PhD.