Ljubljana/finalsystem
From 2007.igem.org
Performance of the Final Functional Systems
Bla bla
Split systems based on heterodimerization
Our results demonstrate that split-ubiquitin and split TEVP systems were activated by the HIV coat protein gp120 which causes dimerization of CD4 and CCR5 receptors. Activation was amplified by the T7 RNAP system connected to luciferase reporter. Figures 4 and 5 show normalized luciferase activities of gp120-stimulated cells, nonstimulated and control cells, respectively in the split ubiquitin (Fig. 4) and split TEVP (Fig.5) system.

Fig. 5. Viral gp120 activates the split TEV protease system. Cells were transfected with CMV–CD4–TEVC, CMV–CCR5–TEVN, myristoylation signal-TEVprotease site–T7 protease, pT7-fLuc and CMV-rLuc reporter plasmids. 12 hours after transfection cells were activated with gp120 protein for additional 12 hours. Cells were lysed and luciferase activity was measured. The results were normalized for transfection efficiency and cell number using rLuc readings. Addition of gp120 activated split TEV protease system as higher amounts of T7promoter luciferase were measured. Increased luminiscence was also observed in control cells, probably due to the same reason as in split ubiquitin experiment.

Fig. 5. Viral gp120 activates the split TEV protease system. Cells were transfected with CMV–CD4–TEVC, CMV–CCR5–TEVN, myristoylation signal-TEVprotease site–T7 protease, pT7-fLuc and CMV-rLuc reporter plasmids. 12 hours after transfection cells were activated with gp120 protein for additional 12 hours. Cells were lysed and luciferase activity was measured. The results were normalized for transfection efficiency and cell number using rLuc readings. Addition of gp120 activated split TEV protease system as higher amounts of T7promoter luciferase were measured. Increased luminiscence was also observed in control cells, probably due to the same reason as in split ubiquitin experiment. T-test: p<0,1, *.T-test: p<0,005, ****.
System based on the activation by HIV protease activity
he process of system activation was observed using fluorescent confocal microscopy. Cells were transfected with CMV–CD4–HIVprotease cleavage site–mCherry–NLS and optionally cotransfected with HIV protease on pNL4.3 plasmid. In cells without HIV protease fluorescence was detected at the plasma membrane (Fig. 8A), whereas in HIV protease cotransfected cells, fluorescent protein mCherry was released from the membrane and translocated to the nucleus (Fig. 8B). Similar results were obtained when mCherry was substituted with T7 RNA polymerase and additional construct pT7–mCerulean was transfected to detect the T7 RNA polymerase activity. In cells with HIV protease cotransfection, mCerulean was detected in cytosol (Fig. 9B), but no fluorescence at all was observed in the control experiment without of the HIV protease activity (Fig. 9A).
The system, using myristoylation signal instead of CD4 to anchor the T7 RNA polymerase to the membrane was also tested and we obtained similar results (data not shown).
We can conclude that in the presence of HIV protease, T7 RNA polymerase is released from the membrane and translocates to the nuceleus where it transcribes genes under the T7 promoter.
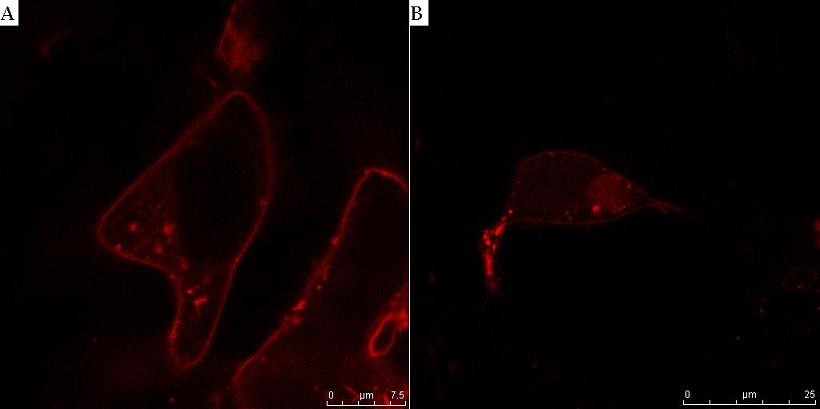
Fig. 8. HIV protease cleaves the linker between the membrane anchor and mCherry. HEK293T cells were transfected with CMV-CD4-HIVprotease cleavage site-mCherry-NLS (A) and additionally with HIV protease plasmid pNL4.3 (B). In cells without HIV protease, membrane localization of mCherry reporter protein was observed (A), but in the presence of HIV protease mCherry-NLS was released to cytosol and translocated to nucleus (B).

Fig. 9. T7 RNA polymerase is activated by the cleavage of the linker on the membrane anchor by HIV protease. HEK293T cells were transfected with CMV-CD4-HIVprotease cleavage site-T7 RNA polymerase-NLS, reporter plasmid T7-mCerulean (A) and additionally with HIV protease plasmid pNL4.3 (B). HIV protease cut off T7 RNA polymerase from the membrane, thus activating it. Consequently mCerulean under the T7 promoter was transcribed and detected in cytosol (B). In the absence of protease the system remained inactive and no fluorescence was observed (A).
Effector protein – caspase-3
Instead of reporter proteins effector proteins such as caspase-3 were used, which are close to the real application. Caspase-3, a cysteine peptidase, which is the main executioner of the apoptosis. We have selected caspase-3 since we wanted to induce apoptosis in HIV-infected cells so that HIV could not replicate and infect the surrounding cells. Apoptosis prevents any inflammation to the neighboring cells and provides for a "clean removal" of apoptotic cells.
Two constructs were prepared for testing the caspase-3 activity: CMV-Cas3 and pT7-Cas3. The first construct was used to test if the caspase-3 is active in transfected cells. The second construct with caspase-3 under the T7 promoter was used as our main effector. Caspase-3 will be transcribed only when active T7 RNA polymerase is present in cytosol or nucleus. This should happen when our devices are activated by HIV virus-induced receptor heterodimerization or by HIV protease, expressed in HIV infected cells.
ELISA was used to determine the amount of caspase-3 in cells – this indicates the apoptotic potential of our system. We wanted to show that amount of caspase-3 in cells infected with HIV protease is greater than in non-infected cells (Fig. 10). The amount of caspase-3 had to be carefully monitored since the expression of caspase-3 leads to apoptosis and killing the cells, which we then can not detect anymore. So, timing of ELISA tests was therefore very important.

Fig. 10. Caspase-3 expression is enchanced in HIV-protease transfected cells. HeLa cells were transfected with effector gene pT7-Cas3 alone or with effector gene and components of system based on activation by HIV protease activity: CD4 membrane receptor with bound T7 RNA polymerase via HIV protease cleavage site and HIV protease in pNL4.3 vector. ELISA test was made with antibodies against human caspase-3. Significant amount of caspase-3 was detected in nontransfected cells and in cells, transfected with pT7-Cas3 as well. Since ELISA can also detect inactive and endogenous caspase-3 and since certain amount of cells is always in apoptosis, we did not expect extremely low values in these two samples. Significantly elevated levels of caspase-3 were detected in cells, transfected with our system components, meaning that more caspase-3 is transcribed and activated cells after HIV protease cuts off T7 RNA polymerase from transmembrane receptor. In terms of our initial idea this means that HIV entry and HIV protease activity in infected cells could trigger apoptosis and thus prevent replication of virus. T-test: p<0,05, **; p<0,01, ***.
Flow cytometry using annexin V staining was used for detection of amount of apoptotic cells as a consequence of caspase 3 activity after system activation. We have transfected cells with components of our systems (that have been previously proved as functional) with caspase-3 under T7 promoter as an effector gene. As a positive control we used cells, transfected with CMV-Cas3 construct. Activation of apoptosis varies between different cell lines and also the efficiency between different inducers varies significantly. Although some apoptosis was detected still an additional optimization is needed to obtain reliable results.
Instead of caspase-3, which triggers apoptosis in HIV infected cells we could use effector proteins that would act as antiviral agents. Good example is interferon-β. Interferons are very important in antiviral defense. They are usually secreted as a consequence of elevated levels of dsRNA in cells. Its effects include prevention of viral replication and stimulation of the immune response. In extreme cases it can also trigger apoptosis, probably trough increased production of p53 gene product.
We have not directly tested the activity of human interferon-β in HIV-infected cells but have deposited BioBricks containing interferon-β, pT7-interferon-β and CMV-interferon-β to the Registry.