Austin Day Notebook
From 2007.igem.org
My Construction Files
My Sequencing Files
AustinDay 16:50, 3 June 2007 (EDT)
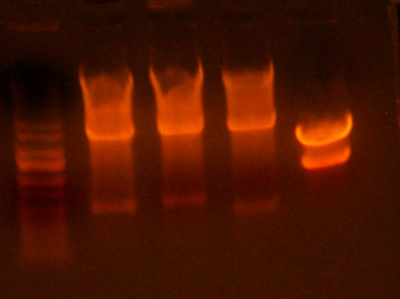
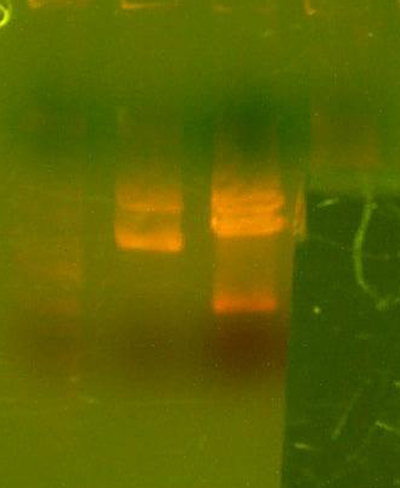
- Made an EcoRI / XhoI digest of the synthetic genes and a digest of EcoRI and XhoI of the 9145 plasmid.
- Here's the gel. Very pretty.
- Took the small fragment from the synthetic genes and the larger fragment from the 9145 plasmid and ligated them together.
- I think I'm going to name these new parts: I716053, I716054, I716055. These names came to me in a dream.
- I grew up 2 colonies of the homogenized RBS pools. (The Kan and the Cmr pools)
Austinday 15:43, 2 June 2007 (PDT)
- Okay, so since the 1121 part was bad, chris found a PCR product of it. I'm going to give that a try and see if it works.
- I did the ligation and transformation for those. I named the parts Bad0001 (kan) and Bad0002 (Cmr). Plated and cooking for tomorrow.
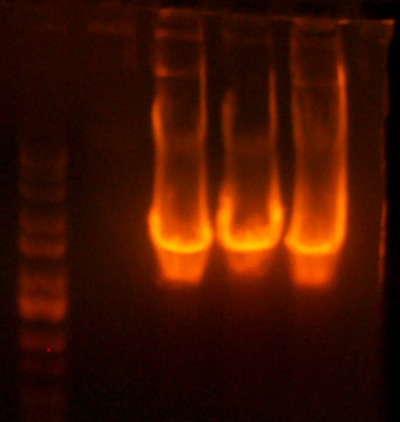
- Made digest of BglII and XhoI of the 3X concentrated 9145 plasmid miniprep. The total digest was equal to that of 14 regular digests. The gel is below:
- Digested the DNA 2.0 plasmids with EcoR1 and XhoI. But...the gel looks funny. I may not have allowed enough time for the lyophilized DNA to redissolve... That's my best guess at least.
- I grew up some colonies of the DNA 2.0 parts so I'll give it another try tomorrow. (Chris told me that there is probably too much DNA and that if I ran it longer, it might have cleared up. You can kind of see that there might be a second band...sort of.)
Austinday 22:59, 30 May 2007 (PDT)
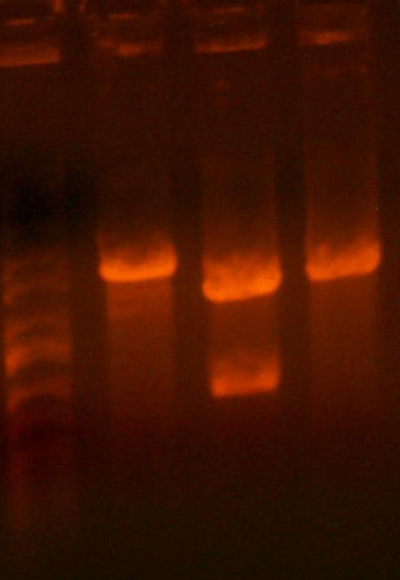
- Okay, I repeated the digestion from yesterday, but got the same results. Here's the gel.
- This time, I let it digest for well over an hour and mixed it half way through. I also ran the gel at a lower voltage than usual.
- The bands for the 1122 lane was noticeably cleaner, but the 1121 lane was uncut.
- I cut out the smaller band from the 1122 lane(because it's probably a higher yield than the previous gel), as well as the 1128 band. I figured that because the previous gel didn't cut completely, and because the 1128 pool smaller bands would have already ran off the gel, this second gel would have a more pure band than the first. I also cut out the region of the 1121 lane where the smaller band should be, although it looks as if nothing is there... I guess we'll have to remake that 1121 part.
Austinday 16:53, 29 May 2007 (PDT)
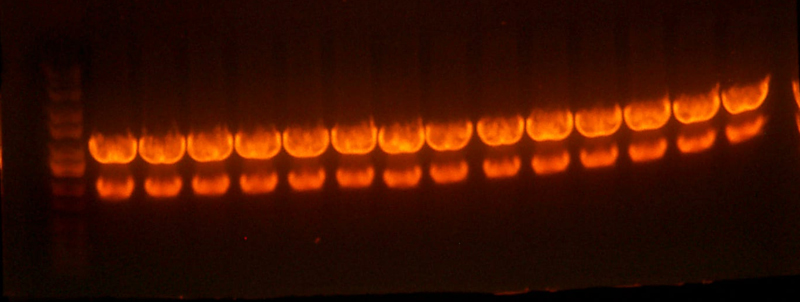
- Sequencing of the pBca1101-Bca1128 RBS library resulted in 2 duplicate RBS's. #24 was identical to #3, and #5 was identical to #2 so I threw out #3 and #5.
- I pooled together the bca1128 rbs's and the 1106 A and B rbs's as well as one other 1128 A rbs Chris had.
- I digested those with EcoR1 and BglII to create the vectors to insert the antibiotic markers. After digestion, the pool looked good so I cut it out right away. The 1121 and 1122 (CmR and Kan) digestions didn't look correct.
- The lanes are: Marker (Barely came out), 1121, 1122, and rbs pool (cut out)
- 1121 has the right number of bands, but I can't exactly tell if the size of the smaller band is correct.
- 1122 has the wrong number of bands. I'm not that experienced with picking out when a band is undigested plasmid, but even if that were the case for the third band, the smallest band looks much too small.
- Anyway, I cut out the smallest bands from 1121 and 1122 and stored them for tomorrow. Maybe Chris will have something to add to my gelatinous adventures.
198.128.27.101 13:55, 25 May 2007 (PDT)
- Preliminary ranking of RBS's picked from the library based on culture colors: (Bca1101 - Bca1128's in TG1)
Stronger --> Weaker
24, 1, 10, 18, 2, 3, 5, 9, 19, 4, 12, 11, 20, 21, 16, 13, 17, 7, 6, 15, 22, 14, 23
The first 7 are significantly more red than the rest.
# 11 was lost due to a tragic mini prepping accident. May it rest in peace.
AustinDay 16:38, 3 June 2007 (EDT)
- Time to start some awesomeness.
- Biobrick numbers: 051-100 Austin Day
- The next few entries are cut and pasted from my Arkin Wiki because I started making entries on that before this wiki was set up.