Duke/Projects/efatf
From 2007.igem.org
Contents |
Electric field-activated transcription factor
Background and Motivation
The λcI repressor (BBa_C0051) is an excellent candidate for screening mutants for a variety of reasons. It is extremely well characterized as a model transcription factor. From its use in its natural settings, we can see that, depending on where the promotor site is located in relation to its target protein, the λcI repressor can act both as a transcriptional activator or repressor (which will be useful in designing a screen). Likewise, the λcI repressor contains a six amino acid sequence (Ser1 Thr2 Lys3 Lys4 Lys5 Pro6) at its extreme N-terminus, known as the 'flexible arm' due to its flexibility apparent in x-ray cryptography, that is found to be essential to promotor binding. Particularly, single mutation and deletion studies have that the Lysines are the most essential, dramatically lowing binding affinity or abolishing binding altogether if removed. We therefore target this region in our mutation studies, the size of which has now shrunk exponentially as a result of the brevity of this extremely important sequence.
Parts and setup
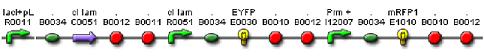
We constructed a genetic circuit as an apparatus to test whether the mutant λcI repressors would bind to DNA or not under a variety of conditions. We combined [http://partsregistry.org/Part:BBa_I5010 BBa_I5010] and [http://partsregistry.org/Part:BBa_I6036 BBa_I6036] with [http://partsregistry.org/Part:BBa_I12007 BBa_I12007] and [http://partsregistry.org/Part:BBa_I13507 BBa_I13507] to yield [http://partsregistry.org/Part:BBa_S03819 BBa_S03819], shown below, on an on an Amp resistant plasmid.
BBa_S03819 contains the λcI promotor (I12007) and repressor (R0051) sequences, nearly identical parts derived from phage λ, which give the λcI repressor protein the ability to activate or repress transcription depending on where the two sequences are placed in relation to the target protein. Thus, BBa_S03819 turns bright RED when the λcI repressor is able bound to DNA (can initiate transcription at I12007 and represses transcription at R0051) and glows YELLOW when λcI repressor is unbound (does not repress transcription at R0051 and does not activate transcription at I12007).
BBa_S03819 was then PCR'd with primers 5’-ccttggatccatAAAGAGGAGAAATACTAGATGNNNNNNNNNAAGAAACCATTAACACAAGAGCAG-3’ or 5’-ccttggatccatAAAGAGGAGAAATACTAGATGNNNNNNNNNNNNNNNCCATTAACACAAGAGCAG-3’, primers with a BamHI restriction sites with random nucleic acid N sections to yield mutants of either the first three amino acids of the flexible arm or the first five amino acids of the flexible arm, respectively, with 5’-aaggggatccgtgctcagtatcttgttatcc-3’; the PCR product was then a linear library of BBa_S03819 mutated at λcI coding region. The three amino acid mutants (called in our lab 'mut3') and the five amino acid mutants ('mut5') were then digested with BamHI, ligated, transformed, and screened for DNA binding under a variety of conditions.
Results
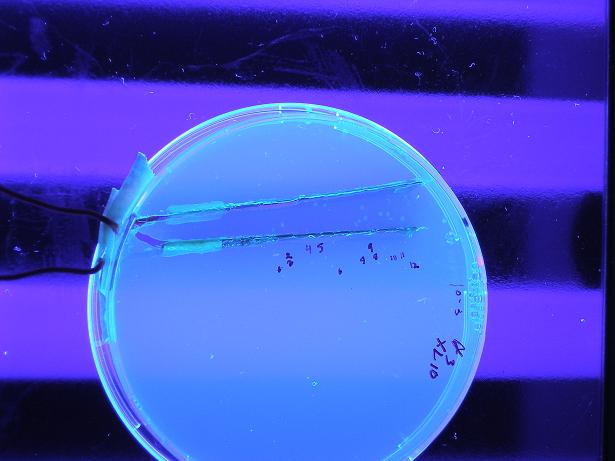
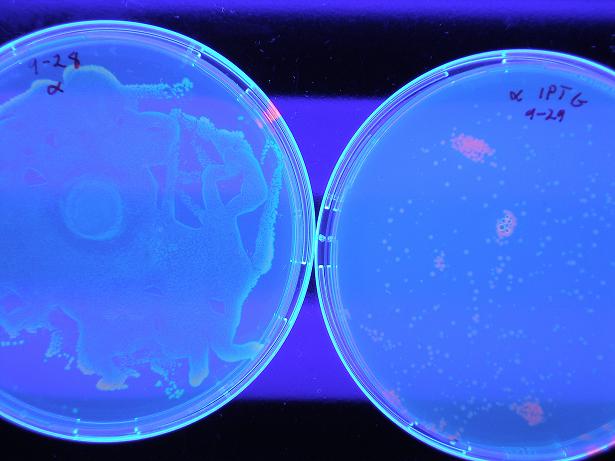
Below shows BBa_S03819 (alpha) both with wild-type λcI proteins unbound to DNA (not present because they are not induced with IPTG) and bound to DNA (wild type λcI proteins are induced with IPTG and present). The red fluorescence along the edges of the uninduced λcI are a result of individual bacterial colonies on the plate with naturally low LacI concentrations, so there exists λcI in the cell; this is not a concern because in screening of the mutants the λcI is always induced (grown on plates with 0.4 uM IPTG), so whether or not the present mutant λcI's bind to DNA is the only screen. Therefore, it is clear the screen works as desired, by glowing red with bound λcI and yellow (which looks green) with unbound λcI.
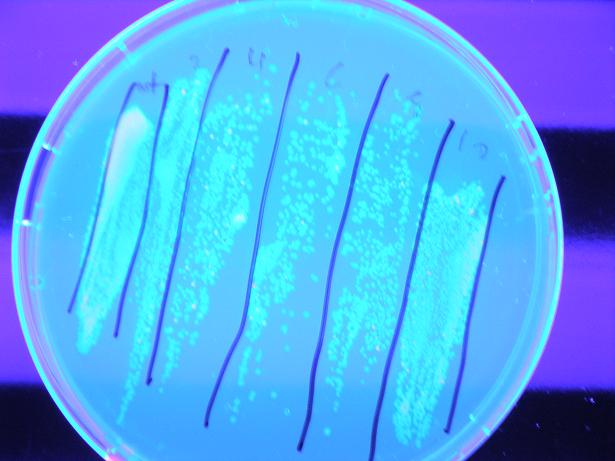
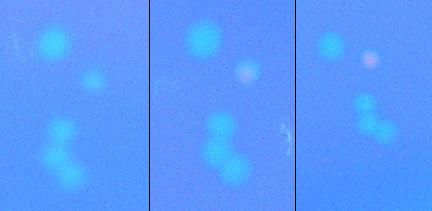
We then attempted to test the apparatus for screening ability before concerning ourselves with the ability to apply large electric fields to bacteria; we did this by attempting to isolate a λcI which at 37C does not bind to DNA, but when heated to 45C would initiate transcription. We plated a plate of alpha_mut3 on a hot-plate set to 45C. Below is a cluster of colonies from the plate at the start of the heating, at 2 hours and at 3.5 hours. It is clear that the bacteria has a λcI which does not preferentially bind to DNA, but upon heating to 45C gains the free energy required to bind to DNA and initiate transcription. These colonies are currently being cultivated so they can be sequenced and tested to determine protein production rates vs. temperature, with results to be presented at our iGEM presentation.
Electric fields were then applied to colonies of bacteria with stainless steel electrodes coated in epoxy (to prevent conductance) attached to a function generator and placed 1 cm apart in a agar plate.
Preliminary tests with fields of 600V(peak-peak)/m at 1kHz, 10kHz, 100kHz, or 1MHz yielded no results (it wouldn't even kill the bacteria!). The heavy machinery was then brought in.
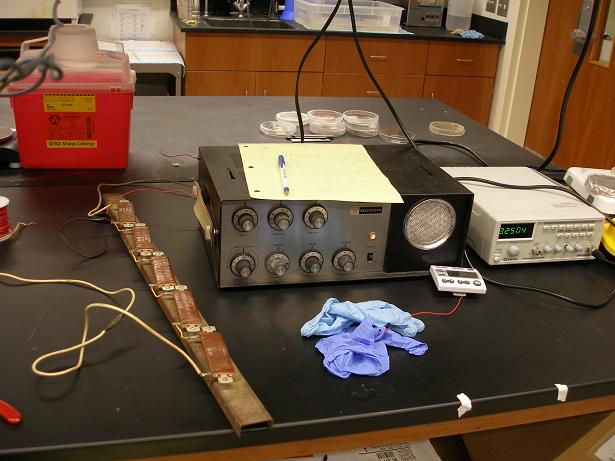
The device above is a Bogen Model M06A vacuum tube audio amplifier. Originally designed to send low voltage audio signals to high voltage speakers, this device generously on loan from Dr. Rhett George is perfect to transform low AC voltage signals into extremely high voltage signals as a result of the conditions necessary to operate vacuum tubes. It took our puny function generator signal (max voltage output 6Vp-p) and output 200Vp-p signals. Below is a picture of bacteria at various output voltage settings at 1kHz in the device, showing that the bacteria can survive in these settings, but probably not much more (and are probably becoming electrocompetent!).
We are now screening mutant colonies in fields of 20kV/m at 1.065 kHz. As we recently acquired this hardware we are screening for electric field-activated proteins at lightning pace to present our findings during our presentation.