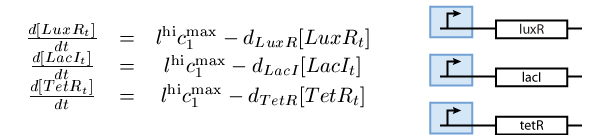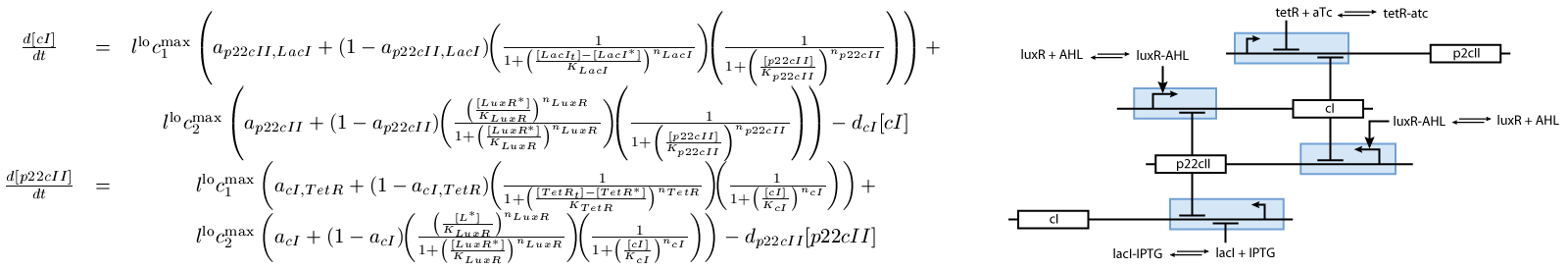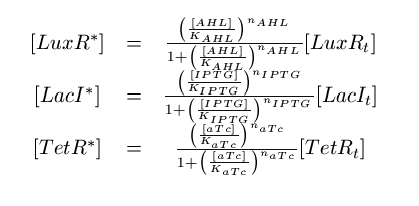ETHZ/Engineering
From 2007.igem.org
(→.:: System Model ::.) |
(→.:: System Model ::.) |
||
| Line 164: | Line 164: | ||
| Ref: [6] (Ratio of association/dissociation) <br> The values of the second bullet are what I've got using the data in the same reference, only considering the systems ''with'' dimerization ([[User:Uhrm|Uhrm]] 09:13, 5 October 2007 (EDT)) | | Ref: [6] (Ratio of association/dissociation) <br> The values of the second bullet are what I've got using the data in the same reference, only considering the systems ''with'' dimerization ([[User:Uhrm|Uhrm]] 09:13, 5 October 2007 (EDT)) | ||
|- | |- | ||
| - | | K<sub>AHL | + | | K<sub>AHL</sub> |
| | | | ||
* 0.009 [mM/s] - 0.1 [mM/s] | * 0.009 [mM/s] - 0.1 [mM/s] | ||
Revision as of 18:28, 7 October 2007

.:: Introduction ::.
In order to understand if it is possible to create the learning system that we wanted, we had to run some initial simulations, to see if we could reach the desirable steady states. After creating a basic framework on which to work on, we refined the parameters by searching the available literatures. In the next, we are presented the coupled differential equations that model our system, their parameters and the values that we picked, the results of our simulations, and lastly, we provide our references. For an introduction to system modeling in synthetic biology, please read our modeling tutorial here.
.:: System Model ::.
Based on [1], we modeled the biological system with differential equations. According to what presented in the Biology Perspective, our system is composed of three subsystems:
- A subsystem of constitutively produced proteins (see Fig. 1),
- The learning part (see Fig. 2), and
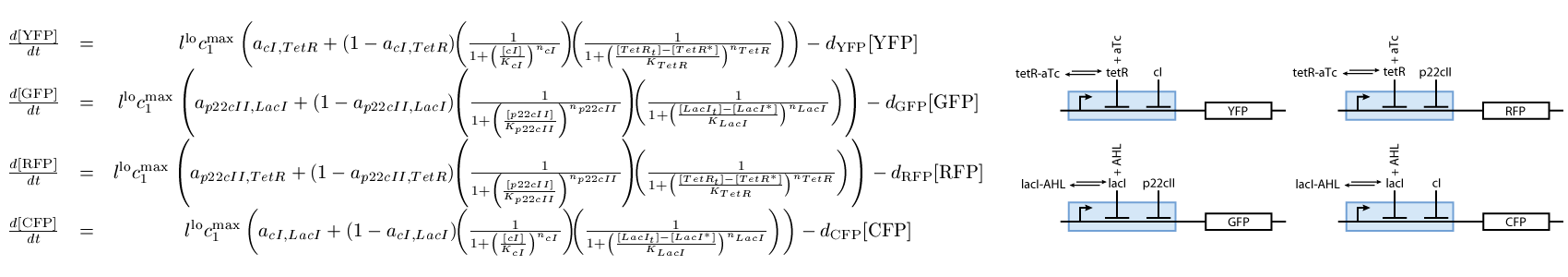
- The reporting subsystem (see Fig. 3).
The second subsystem is the main part of the biological model. This subsystem stores the information concerned the learned chemical, and drives the production of the appropriate reporter, during the recognition phase. It is actually a toggle switch, that reaches a steady state depending on the chemical that it is exposed to (see Fig. 2):
The third subsystem reports the state that our system is, during the different phases of learning and recognition. During the learning phase, this subsystem reports which chemical the cells are exposed to, and during the recognition phase, it reports if the cells recognize the chemicals that they are currently exposed to (see Fig. 3):
Note that the three constitutively produced proteins LacI, TetR and LuxR exist in two different forms; as free proteins and in complexes they build with IPTG, aTc and AHL, respectively. We need to model this complex-forming procedure, with another set of differential equations (Fig. 4):
In order to have meaningful results from our simulations, we browsed through the literature in order to find appropriate values for our parameters. We reduced our parameter space by joining parameters together, and we gave reasonable estimates, for the values that could not be extracted from available publications. Since this is a hard part that every team has to face, we present the table with the chosen parameters below:
| Parameter | Value | Description | Comments |
|---|---|---|---|
| c1max | 0.01 [mM/h] | max. transcription rate of constitutive promoter (per gene) | promoter no. J23105; Reference: Estimate |
| c2max | 0.01 [mM/h] | max. transcription rate of luxR-activated promoter (per gene) | Reference: Estimate |
| lhi | 25 | high-copy plasmid number | Reference: Estimate |
| llo | 5 | low-copy plasmid number | Reference: Estimate |
| ap22cII,LacI | 0.1 - 0.2 | basic production of p22cII/LacI-inhibited genes | Reference: Conclusions after discussion |
| ap22cII | 0.1 - 0.2 | basic production of p22cII-inhibited genes | Reference: Conclusions after discussion |
| acI,TetR | 0.1 - 0.2 | basic production of cI/TetR-inhibited genes | Reference: Conclusions after discussion |
| acI | 0.1 - 0.2 | basic production of cI-inhibited genes | Reference: Conclusions after discussion |
| ap22cII,TetR | 0.1 - 0.2 | basic production of p22cII/TetR-inhibited genes | Reference: Conclusions after discussion |
| acI,LacI | 0.1 - 0.2 | basic production of cI/TetR-inhibited genes | Reference: Conclusions after discussion |
| dLacI | 2.31e-3 [pro sec] | degradation of lacI | Ref. [10] |
| dTetR | 1e-5 [pro sec]/2.31e-3 [pro sec] | degradation of TetR | Ref. [9]/ Ref. [10] |
| dLuxR | 1e-3 - 1e-4 [per sec] | degradation of LuxR | Ref: [6] |
| dcI | 7e-4 [per sec] | degradation of cI | Ref. [7] |
| dp22cII | degradation of p22cII | ||
| dYFP | 6.3e-3 [per min] | degradation of YFP | suppl. mat. to Ref. [8] corresponding to a half life of 110min |
| dGFP | 6.3e-3 [per min] | degradation of GFP | in analogy to YFP |
| dRFP | 6.3e-3 [per min] | degradation of RFP | in analogy to YFP |
| dCFP | 6.3e-3 [per min] | degradation of CFP | in analogy to YFP |
| KLacI | 1.3e-3 - 2e-3 [mM/h] | lacI repressor dissociation constant | lower value is from Ref. [2], higher value is from Ref. [5] |
| KIPTG | 1.5e-10 [mM/h] | IPTG-lacI repressor dissociation constant | Ref. [5] |
| KTetR | 5.6 (+-2) [nM-1] | tetR repressor dissociation constant | Ref. [1] Note: This is for heptameric operator. |
| KaTc | 1120 (+-400) [nM-1] | aTc-tetR repressor dissociation constant | Ref. [1] and Ref. [3]. According to Ref. [3], ratio between tetR repressor dissociation constant and aTc-tetR repressor dissociation constant is 0.4/80. |
| KLuxR |
| luxR activator dissociation constant | Ref: [6] (Ratio of association/dissociation) The values of the second bullet are what I've got using the data in the same reference, only considering the systems with dimerization (Uhrm 09:13, 5 October 2007 (EDT)) |
| KAHL |
| AHL-luxR activator dissociation constant | Ref: [6] (Ratio of association/dissociation) The values of the second bullet are what I've got using the data in the same reference (Uhrm 07:25, 5 October 2007 (EDT)) |
| KcI | 2e-3 [mM/h] | cI repressor dissociation constant | Ref. [5] |
| Kp22cII | p22cII repressor dissociation constant | ||
| nLacI | 1 | lacI repressor Hill cooperativity | Ref. [5] |
| nIPTG</sub> | 2 | IPTG-lacI repressor Hill cooperativity | Ref. [5] |
| nTetR | 3 | tetR repressor Hill cooperativity | Ref. [3] |
| naTc | 2 (1.5-2.5) | aTc-tetR repressor Hill cooperativity | Ref. [3] |
| nLuxR | 2 | luxR activator Hill cooperativity | Ref: [6] |
| nAHL | 1 | AHL-luxR activator Hill cooperativity | Ref. [3] |
| ncI | 1.9 | cI repressor Hill cooperativity | Ref. [5] |
| np22cII | p22cII repressor Hill cooperativity |
.:: Simulations ::.
.:: References ::.
[1] This book
[2] A synthetic time-delay circuit in mammalian cells and mice (http://www.pnas.org/cgi/content/abstract/104/8/2643)
[3] Detailed map of a cis-regulatory input function (http://www.pnas.org/cgi/content/full/100/13/7702?ck=nck)
[4] Parameter Estimation for two synthetic gene networks (http://ieeexplore.ieee.org/iel5/9711/30654/01416417.pdf)
[5] Supplementary on-line information for "A Synthetic gene-metabolic oscillator" (no link)
[6] Genetic network driven control of PHBV copolymer composition (http://doi:10.1016/j.jbiotec.2005.08.030)