Samantha Liang Notebook
From 2007.igem.org
Revision as of 23:03, 19 June 2007 by Samanthaliang (Talk | contribs)
My Construction Files
My Sequencing Files
My Biobricks
Samanthaliang 15:26, 19 June 2007 (EDT)
- Grow up one colony of 214 and 215 in Lefty, and 210 and 209 in Righty
- mini 212 and sequence it again
- actually didn't mini the 223s and do an analytic gel to see if any are good because I did a colony PCR and they came out promising
- Plates of the new cassette+toxins looked great
- Colony PCR on the cassette+toxins
Samanthaliang 14:54, 18 June 2007 (EDT)
- grew up 2 clones of 212 that i subcloned again
- sequencing came back for the cassette+toxins and only 220, 221, 222 are good.
- Subcloning all the others again with more DNA in the digest.
- Also, will grow up some more colonies of 223 to screen tomorrow because it uses the same insert as 222 and the same backbones as 220, 221, so it is possible that some of the colonies may be correct.
- Also, will try out the 123 cloning method on 216 and 217 (218 and 219 too). First step is to transform the inserts and backbones parent plasmids into the special strains. Transformed 214 and 215 into "Lefty" competent cells, then 209 and 210 into "Righty" competent cells.
Samanthaliang 18:10, 15 June 2007 (EDT)
- miniprep 216-223 clones
- test digest 216-223 clones
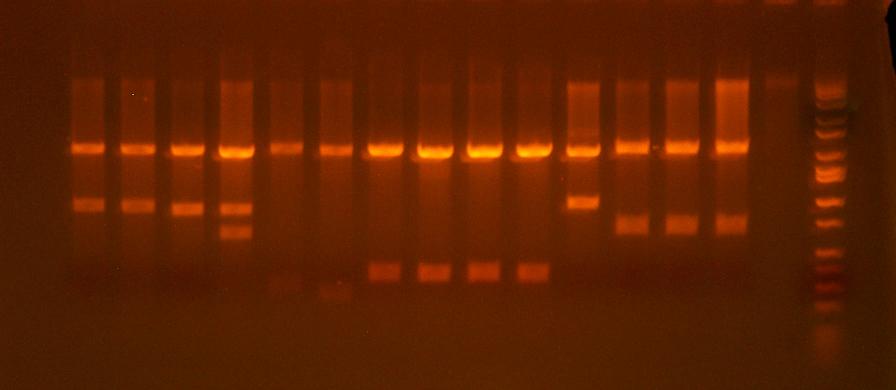
from left to right:
I716216-1, I716216-2, I716217-1, I716218-2, I716218, I716219,I716220-1, I716220-2, I716221-1, I716222-1, I716222-2, I716223-1, I716223-2, blank, ladder
- clones that pass the test digest: 220-1, 220-2, 221-1, 221-2, 222-1
- I'm also clone saving and sequencing 216-1 and 223-1 just because I want to see what's up with them. 216 and 217s don't show the pattern of either the desired product or either vector. 223-1 is a maybe, it may be missing the cassette. 218 and 219 both failed miserably, but that was to be expected.
- accidentally grew up 212, 218 and 219 on plain LB plates with no Carb. Arg. Religate and transform again from previous digests
- If the 218 and 219 plates come out poorly again, will have to redo the I716210 digest (BglII/BglI), should have enough of the cassette though.
- Read that RNA destabilization article
Samanthaliang 15:28, 14 June 2007 (EDT)
- sequencing for the GTG start Cre was same as before - still has an ATG start - I can't really figure out what is going wrong here. I definitely used dpnI to digest the parent vector because I remember having to go upstairs to the Keasling lab to get hooked up with that enzyme since we were out of it downstairs. Also, both times I did it, there were a lot of colonies on the plate, and there would have been none if it were the parent vector since I used dpnI. It must be some problem with the PCR then - is it self-correcting? I double checked the oligo and it definitely has a GTG start. I guess I'll try to make it one more time, but if it doesn't work this time then I might have to try Quick Change if it's worth having this part.
- Grow up colonies for cassettes+toxins (216-223)- the plates are alright. except for the ones with BamHI, those plates only had one colony each. weird. maybe a problem with the digest since both of them are bad? Will re-ligate and transform today hoping for an easy fix. If the sequencing comes out bad though eventually, I will have to redo the subcloning for 218 and 219 from the start.
- Make the ape files for cassette+toxins
- Read that locks and keys article
- Subclone Cre (GTG) over.
Samanthaliang 18:21, 13 June 2007 (EDT)
- miniprepped 212s - sequenced the clones that came from the plate with more colonies (put cultures in fridge)
- subcloned all the cassettes+toxins (216-223) because sequencing came back good on the cassettes for the clone 2s - 8 different new plasmids!
- put teeth on the centrifuge
Samanthaliang 15:26, 12 June 2007 (EDT)
- grew up 212 clones
- miniprepped 214 and 215 clones and sent them out for sequencing
- clone saved
- sequencing came in and it looks like we have a good clone for every toxin!
- ceaB (209) - clone 2
- BamHI (210)- clone 2
- barnase (211)- clone 1
- BglII - already a miniprep Bca1147
- still missing Cre with a GTG start - is being grown up today
- Cre with a TTG start (213) clone 2 is perfect
- Made construction files for cassette+toxin
- Now in charge of the google spreadsheets - have to invite people and what not
For tomorrow: Once the cassettes have been confirmed for sure, can digest those and the toxins and ligate.
Samanthaliang 19:24, 11 June 2007 (EDT)
- grew up 214 and 215 clones
- Sequenced 209 (clone 1&2), 211 (clone 1&2), and 213 (clone 2) again with G01001 (backwards) because the genes were too big to confirm for sure that they were right, 210 was perfect the first time
- Redid the PCR for 212 (Cre-GTG) because both sequences came back with the ATG start codon still
- Subcloned 212
- Clone saved and recorded on the new Google Spreadsheet
- emailed china team that wanted our oriT knockouts from last year - told them to look in the registry again
Samanthaliang 20:00, 9 June 2007 (EDT)
- Miniprepped 209, 210, 211, 212, 213 and sent them out for sequencing
- Sequencing came in for 207 and 208 - should use clone 1 for both.
Will try to come in on Sunday too to grow up the 214 and 215 cultures - i moved the plates to the fridge for now.
Samanthaliang 17:58, 8 June 2007 (EDT)
- Made a ridiculously large batch of DH10B competent cells
- Miniprepped 207 and 208 clones, clone saved them, and sent them out for sequencing
- Used those preps to subclone I716214 and I716215 (two versions of each)
- Grew up cultures of 209, 210, 211, 212, 213
Samanthaliang 17:58, 7 June 2007 (EDT)
- Chris gave me the wrong template for Cre and I didn't notice until today so I redid those 2 PCRs
- All the other PCRs looked fine on a gel
- Cleaned up Digested and inserted all PCR products into Biobrick form (I716210 through I716113)
- Got sequencing data back and only one of the [rbs][atg] ones came out good, but the [lox][terms]s all came out good - just forget about the [rbs][atg] ones for now. More detail - the ones that failed also had really tiny colonies and were in Bca1101 form with a promoter in front, so it might be translating some nonsense that is toxic to the e.coli - but since I have one of the rbs's already I'm just going to proceed with that
- Picked colonies from pBca9145-I716207-1 and pBca9145-I716208-1
- Austin grew up 10ml of DH10B for me so I can make competent cells tomorrow
Samanthaliang 17:31, 6 June 2007 (EDT)
- miniprepped 201 through 206 and sent them out for sequencing
- put the clones on "clone saver" paper
- subcloned 207 and 208 with all clones (have 2 versions of each just in case)
- PCRed ceaB, BamHI, Barnase, Cre with both alternate stop codons
Samanthaliang 19:20, 5 June 2007 (EDT)
- Grew up 2 colonies each of I716201 through I716206
- Made a lot of construction files and entered things into the registry
- My oligos were ordered today!
- Demonstrated liquid media making and registry entering
- Now will also make an RFP variant as a reporter on how well Cre is working
To do tomorrow:
- subclone 207 and 208
- Run PCRs if they come in the afternoon
- Send things out for sequencing and make -80s
- Fill out and sent those survey forms
Samanthaliang 15:29, 4 June 2007 (EDT)
- Oligos to make biobricks of toxins are not yet in - maybe Wednesday or Thursday?
For cassette between Cre and toxin: [rbs][ATG][Lox][Term][Lox]
- Made [rbs][ATG] variants with different ribosome binding sites
I716201 is Bca1106A.Bca1112
I716202 is Bca1106B.Bca1112
I716203 is Bca1128A.Bca1112
I716204 is Bca1090.Bca1112
- Made [Lox][Term] variants with 2 different terminators (will probably use Bca1092 version)
I716205 is Bca1114.Bca1124
I716206 is Bca1114.Bca1092
- Tomorrow will make [Lox][Term][Lox] part
- Then will put the rbsATG together with LoxTermLox
eek so many construction files and ape files to make.
to do