Paris/Results
From 2007.igem.org
Contents |
Experimental results
dapA deletant growth
We expect the somatic cells to excrete only a limited amount of DAP, it would thus be nice if our auxotroph strain could grow on limited amount of metabolite. Over night culture were diluted 100 time in LB suplemented with different amount of DAP.
DAP excretion by prototroph cells
We want to know if prototroph cells are excreting DAP and in what amount. Instead of doing proper chemical dosages which are expensive, we had the idea to make a biological measurments.
Prototroph MG1655 strain was grown in LB, and the medium was recovered at different stages of the culture and purified. The dapA deletant strain was then grown in the recovered medium. The idea is that comparing the growth curve of this experiment with growth curves of dapA deletant in differently supplemented LB medium, should alow us to determine approimatively the amount of DAP present in the recovered medium.
Survival of dapA deleted strain and Co-culture experiments
If we want our system to be robust, the germ line cells should be able to live as long as possible when deprived of DAP. They may indeed face such a feat within our system if their is not enough somatic cells at a given time.
Our dapA deleted strain was grown over night in LB + 300µM DAP. A culture was then launched diluting 100 time the ON culture in LB (+ Kanamycin). 100µL were plated each hour and the number of clone determined.
We also want to know if the soma of our synthetic organism will be able to feed the germ line. Unfortunatly, this cannot properly be done without the final construct. Nevertheless, we can already check if in similar situations, dapA deleted strain can survive.
Two different co-culture experiments were performed:
- In the first one, the dapA deleted strain is grown with a prototroph strain. In this case, we know for sure that the prototroph cells will take the population over. Nervertheless it is interesting to see if dapA deleted strain survive longer in this condition than if alone and without DAP.
(the experiment has only been done once, and would be worth repeating to confirm the results)
We can draw from this curves two important conclusions:
1. Most of the cells die within the first 2 hours of culture, but a small fraction of the cells are still alive after 8 hours. A part of the dapA deleted cells can thus survive for quite long, which is a good point for us.
2. When in coculture with a prototroph cell, around 100 time more cells have a long term survival. This means that the dapA deleted strain benefits a bit from the prototroph cell presence and its DAP production capacities.
- In the second coculture experiment, the dapA deleted strain is grown with a strain bearing an auxotrophy to another metabolite. This experiment reproduces cross-feeding works done on the yeast by Shou W et al. (Synthetic cooperation in engineered yeast populations, PNAS). In this coculture there is a mutual dependence of each strain for the other. If this works, then we know for sure that dapA deleted strain can be rescued by a strain producing DAP!
Our dapA- strain was grown with a tryptophan operon deleted strain. The culture was plated to check the presence of both strains after 8H. They were both present in concentration above 5*10^4 CFU/Ml. This clearly means that DAP is a good choice as our auxotrophy metabolite !
Recombination frequency measurements
In the section “Design Process”, the question of the recombination frequency has been discussed. G to S differentiation frequency, and thus lox recombination frequency on which it is based, is a key aspect of our system. Indeed, optimal overall differentiation frequency lies somewhere between 0 and 50% per generation.
We would like to have tuneable recombination device in order to find the optimal frequency, which lies somewhere between 0 and 50% of recombination per generation. We should be able to tune the recombination frequency by modulating the Cre recombinase expression. In order to do this, we have cloned Cre under the control of the pBAD promoter. We then wanted to characterize our "Cre generator device", and determine the relation between Cre expression level and recombination frequency.
Three types of experiments were planned and partially performed in this regard:
- Experiments using FX85 strain harbouring lox-KmR-lox casstte: indirectly measuring recombination frequency by plating on Kan plates after Cre expression assays.
- Experiments using FX85 strain harbouring lox-KmR-lox casstte: indirectly measuring recombination frequency by studying growthrates.
- Experiments using the recombination measurement biodevice.
A) Experiments using FX85 strain harbouring lox-KmR-lox casstte: indirectly measuring
We used FX85 strain (provided by Francois-Xavier Barre) carrying the cassette: lox-KmR-lox inserted into its chromosome. We transformed this strain with our pBad-Cre construct (carried by an Ampiciline resistance plasmid: pSB1A2). Inducing of Cre expression with different arabinose levels was performed. The last step is then spreading on selective plates (Amp or Amp+Kan). The ratio between the number of clones on the two types of plates should give us an estimate of the recombination frequency. When we tried to do this, we did not get any clones on the Amp+Kan plates, even without Cre induction. But this doesn't mean that the recombination frequency is 100%. In fact, if recombination frequency is around 50% per generation or higher, no colonies can grow on kan plates, simply because half or more of the cells will die at each generation (those who recombine are not resistant to Kanamycin any more after little time).
This means that our Cre generator is quite leaky and gives quite high recombination rates without induction.
B) Experiments using FX85 strain harbouring lox-KmR-lox casstte: indirectly measuring recombination frequency by studying growthrates.
If recombination frequency is two high in owr experimental system, then no growth can be seen on Ampicilline (plasmid selection antibiotic)+ Kanamycin (screens against recombiants: cells having excised lox-kanR from their genome). We tested an alternative strategy in order to circumvent this problem. In order to approximately determine what recombination rate we had, we performed another experiment. FX85 strain was transformed with our pBad-Cre construct. Directly after transformation, liquid cultures were launched with either Amp or Amp+Kan. 100µl of the cultures were regularly plated on LB+Amp, giving the following growth curves:
C) The third strategy is based on a more direct observation of recombination rate. We have constructed a “Recombination frequency measurement” device. The schematic structure of this genetic construct is as follows:
dapAp characterisation
pBAD characterisation
DGAT cloning and triglyceride synthesis in E. Coli
- TG synthesis experiment
1. We transformed chemically competent E.coli (DH5alpha) with pBluescript SK minus vector (Stratagene) (ampicilline resistance and pLac promoter) baring DGAT gene (pKS::DGAT). In this vector, dgat transcription is induced by IPTG.
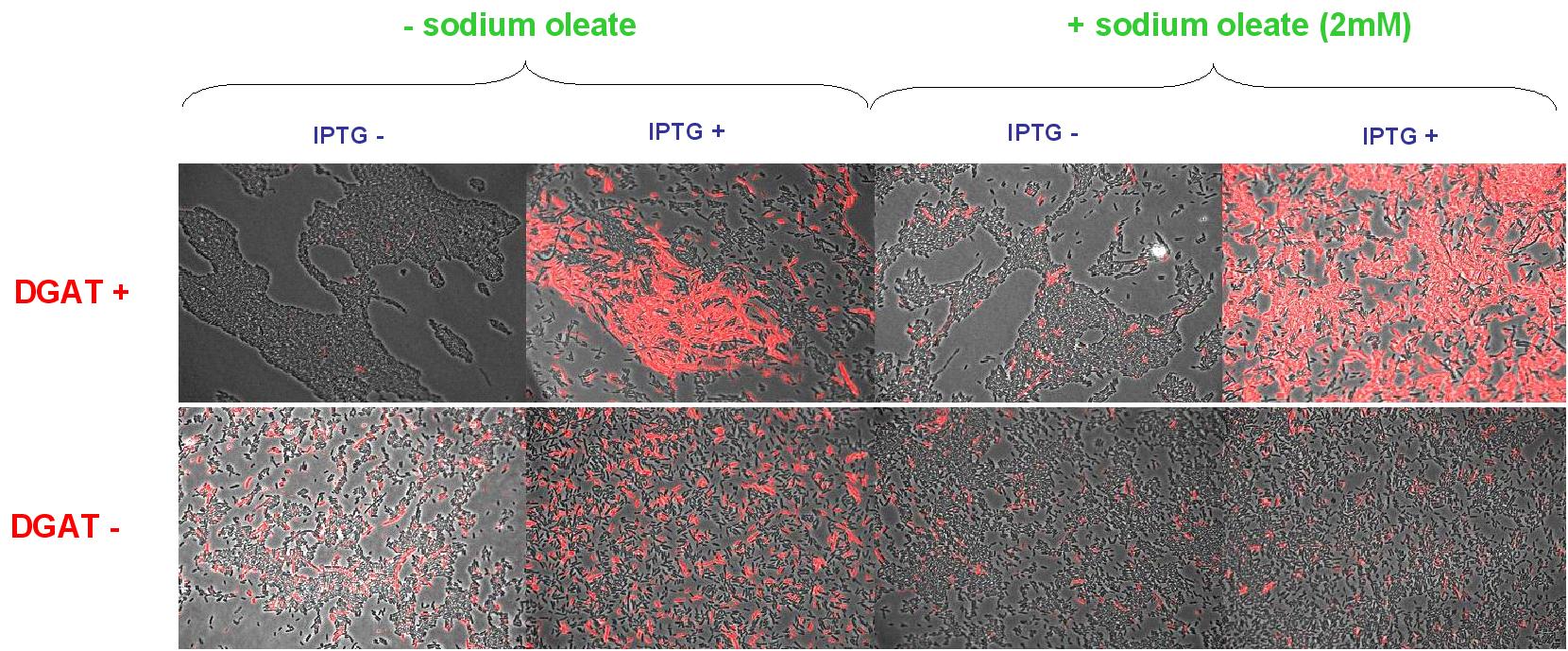
Nile Red fluorescence dye was used, at a concentration of 5µg/mL to monitor lipid inclusions in different conditions of DGAT expression and fatty acid availability:
(0.4mM IPTG induction in LB medium with or without sodium oleate 2mM). Results are shown below.
Line 1 represents E.coli transformed with pKS::DGAT; and Line 2 the negative control (E.coli transformed by part B0015). Columns 1 and 2 are LB medium without sodium oleate supplementation; columns 3 and 4 are LB with sodium oleate supplementation (2mM). Columns 1 and 3 are without IPTG; columns 3 and 4 with IPTG induction (0.4mM).
We can observe lipid inclusion into E.coli transformed by pKS::DGAT with IPTG induction.
No significant difference is seen between the +/- oleate cells.
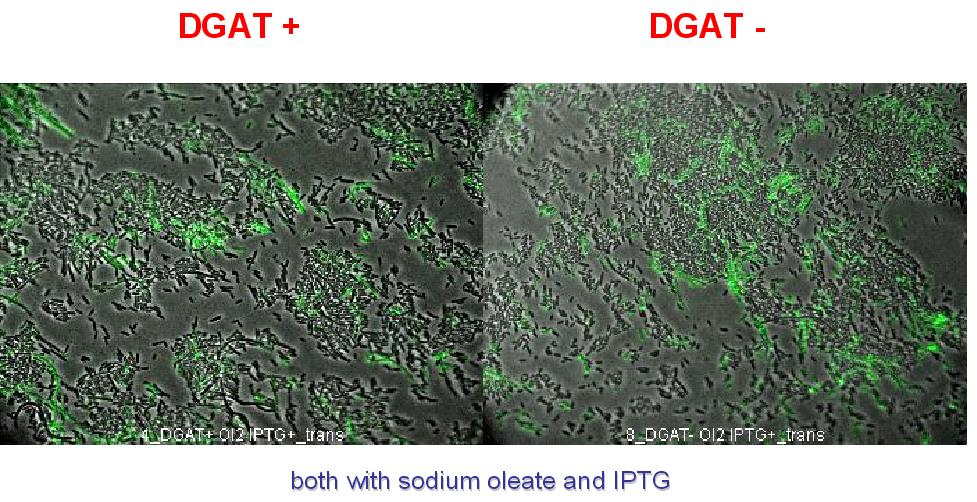
2. To exclude that the fluorescence observed is due to DGAT induced cell death. A cell death marker (green) is used. It can be seen below that cell death is not increased upon DGAT expression.
3. We started creating DGAT biobrick BBa_I718002:
- PCR based mutagenesis was performed to eliminate a PstI site in dgat coding sequence.
- We attempted adding biobrick prefix and suffix sites to dgat but have yet to finish the cloning process :-(