Davidson Missouri W
From 2007.igem.org
Macampbell (Talk | contribs) (→Our Project) |
Macampbell (Talk | contribs) (→Our Successful Project) |
||
| (3 intermediate revisions not shown) | |||
| Line 14: | Line 14: | ||
! style="color: white; background-color: black;"| The Team | ! style="color: white; background-color: black;"| The Team | ||
! style="color: white; background-color: black;" | The Faculty | ! style="color: white; background-color: black;" | The Faculty | ||
| - | ! style="color: white; background-color: black;" | | + | ! style="color: white; background-color: black;" | School Logos |
! style="color: white; background-color: black;" | Group Photo | ! style="color: white; background-color: black;" | Group Photo | ||
|- | |- | ||
| Line 85: | Line 85: | ||
<center> | <center> | ||
| - | =Our Project= | + | =Our Successful Project= |
</center> | </center> | ||
| Line 113: | Line 113: | ||
As a part of iGEM2006, a combined team from Davidson College and Missouri Western State University reconstituted a hin/''hix'' DNA recombination mechanism which exists in nature in ''Salmonella'' as standard biobricks for use in ''E. coli''. The purpose of the 2006 combined team was to provide a proof of concept for a bacterial computer in using this mechanism to solve a variation of The Pancake Problem from Computer Science. This task utilized both biology and mathematics students and faculty from the two institutions. | As a part of iGEM2006, a combined team from Davidson College and Missouri Western State University reconstituted a hin/''hix'' DNA recombination mechanism which exists in nature in ''Salmonella'' as standard biobricks for use in ''E. coli''. The purpose of the 2006 combined team was to provide a proof of concept for a bacterial computer in using this mechanism to solve a variation of The Pancake Problem from Computer Science. This task utilized both biology and mathematics students and faculty from the two institutions. | ||
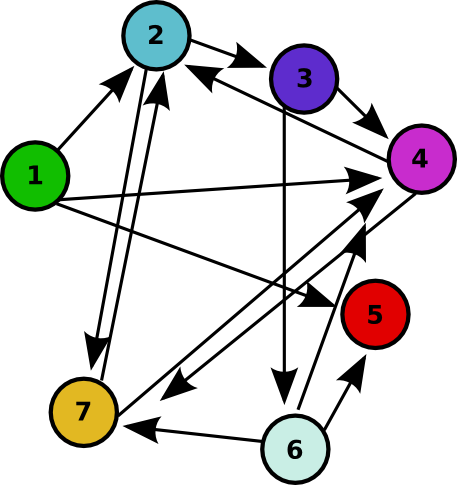
| - | For 2007, we successfully continued our collaboration and our efforts to manipulate ''E. coli'' into mathematics problem solvers as we refine our efforts with the hin/''hix'' mechanism to explore another mathematics problem, the Hamiltonian Path Problem. This problem was the subject of a groundbreaking paper by Adleman in 1994 (see [[Davidson_Missouri_W/Resources_and_Citations | citations]]) where a unique Hamiltonian path was found ''in vitro'' for a particular directed graph on seven nodes. We | + | For 2007, we successfully continued our collaboration and our efforts to manipulate ''E. coli'' into mathematics problem solvers as we refine our efforts with the hin/''hix'' mechanism to explore another mathematics problem, the Hamiltonian Path Problem. This problem was the subject of a groundbreaking paper by Adleman in 1994 (see [[Davidson_Missouri_W/Resources_and_Citations | citations]]) where a unique Hamiltonian path was found ''in vitro'' for a particular directed graph on seven nodes. We were able to use bacterial computers to solve the Hamiltonian path problem ''in vivo''. ([[Davidson Missouri W/Background Information#Why Use Bacteria?|Why use a bacterial computer?]]) |
<br> | <br> | ||
| Line 125: | Line 125: | ||
<br> | <br> | ||
| - | <center> '''A Human Representation of the Adleman Graph.''' | + | <center> '''A Human Representation of the Adleman Graph. (mouse over to see the full effect)''' |
<html> | <html> | ||
Latest revision as of 04:32, 27 October 2007
The Team
| The Team | The Faculty | School Logos | Group Photo |
|---|---|---|---|
| Davidson
Oyinade Adefuye
| |||
| Missouri Western
Jordan Baumgardner
| 
|
Our Successful Project
| In Depth | Overview | ||
|---|---|---|---|
|
Background Information
| Hamiltonian Path Problem
As a part of iGEM2006, a combined team from Davidson College and Missouri Western State University reconstituted a hin/hix DNA recombination mechanism which exists in nature in Salmonella as standard biobricks for use in E. coli. The purpose of the 2006 combined team was to provide a proof of concept for a bacterial computer in using this mechanism to solve a variation of The Pancake Problem from Computer Science. This task utilized both biology and mathematics students and faculty from the two institutions. For 2007, we successfully continued our collaboration and our efforts to manipulate E. coli into mathematics problem solvers as we refine our efforts with the hin/hix mechanism to explore another mathematics problem, the Hamiltonian Path Problem. This problem was the subject of a groundbreaking paper by Adleman in 1994 (see citations) where a unique Hamiltonian path was found in vitro for a particular directed graph on seven nodes. We were able to use bacterial computers to solve the Hamiltonian path problem in vivo. (Why use a bacterial computer?)
Click here for the solution. | ||
<Previous Section | Next Section>