Paris/July 16
From 2007.igem.org
Nicolas C. (Talk | contribs) (→PCRs) |
(→Plasmid pKs::DGAT expression in E. Coli) |
||
| Line 3: | Line 3: | ||
== Plasmid pKs::DGAT expression in E. Coli == | == Plasmid pKs::DGAT expression in E. Coli == | ||
We tried to see TG with NR died on E. Coli cells transfected with pKs::DGAT, an IPTG-inducible promoter. | We tried to see TG with NR died on E. Coli cells transfected with pKs::DGAT, an IPTG-inducible promoter. | ||
| - | We tried different growth media containing more or less oleate, that should in theory increase TG synthesis.<br> | + | We tried different growth media containing more or less oleate (0.5 and 2mM), that should in theory increase TG synthesis. |
| - | [[Image:Paris_EColi_DGAT.jpg]]<br> | + | <br> |
| - | Results : we don't see the | + | [[Image:Paris_EColi_DGAT.jpg]] |
| + | <br> | ||
| + | Results : we don't see the inducible effect of IPTG. We can think that : | ||
* Either the fluorescence without IPTG is due to a leak of the promoter | * Either the fluorescence without IPTG is due to a leak of the promoter | ||
* Either DGAT is not induce in presence of IPTG, and the fluorescence we see is only a background. | * Either DGAT is not induce in presence of IPTG, and the fluorescence we see is only a background. | ||
| + | <br> | ||
| + | |||
| + | We can also wait to see accumulation of TG. Indeed in Acinetobacter, the process is pretty long (2-4 days). It could be the same for E.coli transformed by pKS::DGAT. | ||
== Growth kinetics of w121 strain == | == Growth kinetics of w121 strain == | ||
Revision as of 12:34, 30 July 2007
Contents |
Plasmid pKs::DGAT expression in E. Coli
We tried to see TG with NR died on E. Coli cells transfected with pKs::DGAT, an IPTG-inducible promoter.
We tried different growth media containing more or less oleate (0.5 and 2mM), that should in theory increase TG synthesis.
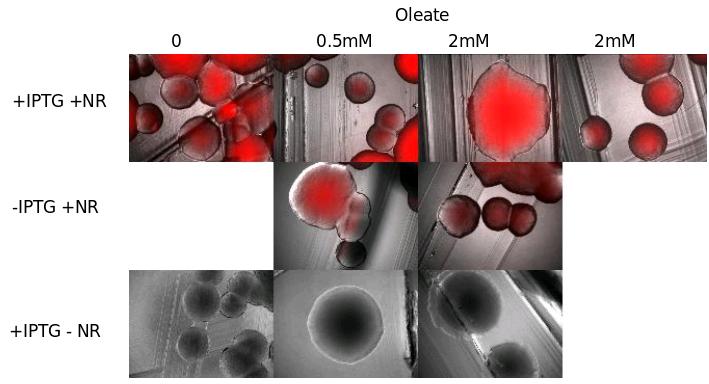
Results : we don't see the inducible effect of IPTG. We can think that :
- Either the fluorescence without IPTG is due to a leak of the promoter
- Either DGAT is not induce in presence of IPTG, and the fluorescence we see is only a background.
We can also wait to see accumulation of TG. Indeed in Acinetobacter, the process is pretty long (2-4 days). It could be the same for E.coli transformed by pKS::DGAT.
Growth kinetics of w121 strain
Results of the previous day : we lost everything because of a crash of the computer :( Sorry Eimad !
Transduction of MG1655 with P1 stock made on w121
- Control (1mL LB MgSO4 30mM; CaCl2 15mM)
- 5µL Phage + 900µL LB (MgSO4 30mM; CaCl2 15mM) + 100µL MG1655 Culture ON
- 50µL Phage + 900µL LB (MgSO4 30mM; CaCl2 15mM) + 100µL MG1655 Culture ON
- 500µL Phage + 500µL LB (MgSO4 30mM; CaCl2 15mM) + 100µL MG1655 Culture ON
=>For the nth time, it is not working : we have only contaminants.
MiniPreps
- I0500 clones 1, 2
- pJ23107 clones 1, 2
PCR purification
- Lox71-FtsZ1
- FtsZ2
- DGAT1
- DGAT2
PCRs
The Lox66-DapAColi PCR did not work... We'll try again with different annealing temperature
Assembly PCRs:
- Lox71-FtsA-FtsZ-1 + FtsZ-2
- DGAT-1 + DGAT-2
The Lox66-DapAColi PCR did not work... so we try a gradient of annealing temperatures
Transformations of Biobricks
As in the protocol given in the registry. Transformation of biobricks :
- BBa_B0030 pSB1A2 : RBS (well 3G plate 1)
- BBa_E0422 pSB1A2 : ECFP (RBS+LVA+Term) (well 11G plate 1)
- BBa_E0241 pSB1A2 : PoPs to GFP converter (well 15c plate 2)
- BBa_E0840 gfp tri-part; strong rbs (well 16E plate 1)
- BBa_J61047 pSB1A2 : Cre ORF (well 8P plate 4)
Transformation in DH5alpha subcloning efficiency
Spread on LB-Amp