ETHZ
From 2007.igem.org
| Line 94: | Line 94: | ||
... this is what George Orwell would have written, were he a synthetic biologist. In the <i>E.coli</i> colonies on petri dishes, all bacteria are equal; except for some special ones. Our project is about modeling and designing these special <i>E.coli</i> that are "more equal" than the rest: they have the ability to be trained, memorize, and recognize their environment, and their story will be presented through this wiki ... | ... this is what George Orwell would have written, were he a synthetic biologist. In the <i>E.coli</i> colonies on petri dishes, all bacteria are equal; except for some special ones. Our project is about modeling and designing these special <i>E.coli</i> that are "more equal" than the rest: they have the ability to be trained, memorize, and recognize their environment, and their story will be presented through this wiki ... | ||
| - | =Motivation= | + | ==Motivation== |
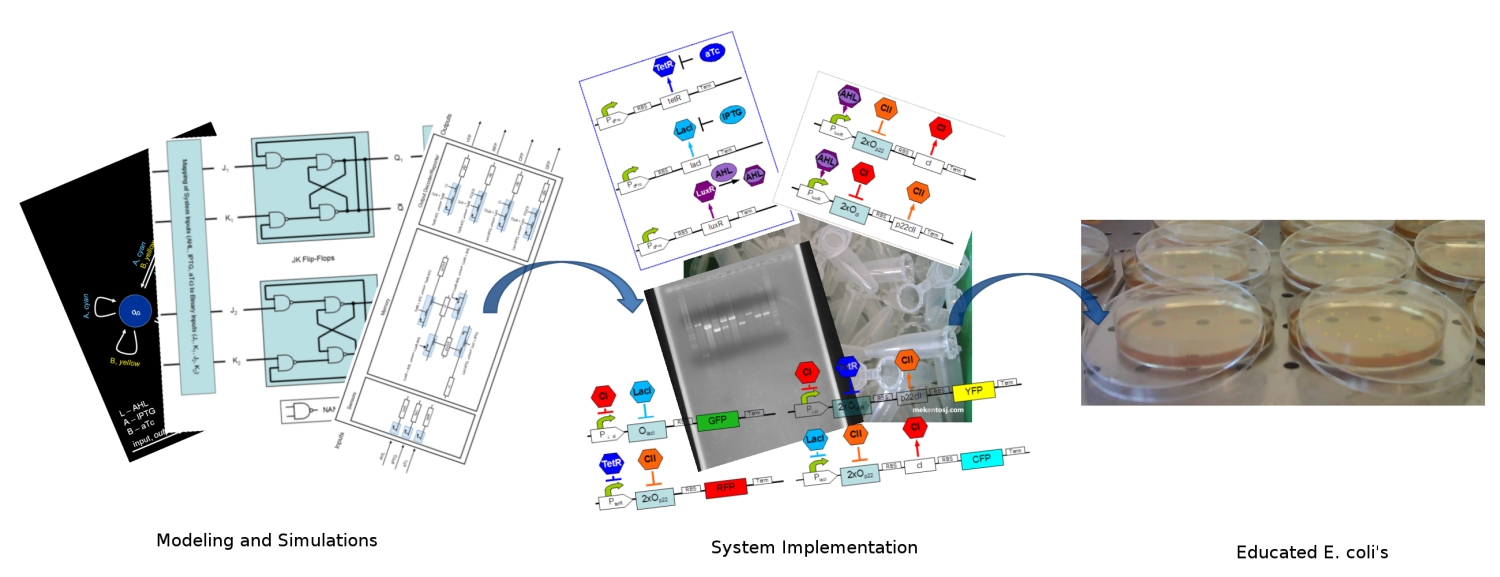
[[Image:ethz_main_pic.jpg|right|thumb|<b>Fig. 1</b>: Artist's approach to the different stages of the development. We started by modeling and simulating the system, we continued by specifying the DNA strands for its implementation, and in the end, our system should report with different fluorescent proteins (image edited)|600px]] | [[Image:ethz_main_pic.jpg|right|thumb|<b>Fig. 1</b>: Artist's approach to the different stages of the development. We started by modeling and simulating the system, we continued by specifying the DNA strands for its implementation, and in the end, our system should report with different fluorescent proteins (image edited)|600px]] | ||
| Line 102: | Line 102: | ||
It is obvious thus, that we are coping with the problem of implementing memory capabilities in bacterial colonies. Our system is a memorizing system, and it also has the ability to understand its environment through a recognition phase. If we assume that the training chemicals are harmful for humans, we can use the developed system to understand whether a particular environment is dangerous for humans or not. In this sense, our system can have applications in the health/safety sector, as is described below: | It is obvious thus, that we are coping with the problem of implementing memory capabilities in bacterial colonies. Our system is a memorizing system, and it also has the ability to understand its environment through a recognition phase. If we assume that the training chemicals are harmful for humans, we can use the developed system to understand whether a particular environment is dangerous for humans or not. In this sense, our system can have applications in the health/safety sector, as is described below: | ||
| - | + | ===Intelligent Biosensors and Self-Adaptation=== | |
We constructed a system capable of sensing different chemicals and producing different fluorescent proteins. Since the cells can be trained to produce one of several specific fluorescent protein types when a certain chemical is present, one can also view those cells as intelligent biosensors, able to change their properties in a training phase. | We constructed a system capable of sensing different chemicals and producing different fluorescent proteins. Since the cells can be trained to produce one of several specific fluorescent protein types when a certain chemical is present, one can also view those cells as intelligent biosensors, able to change their properties in a training phase. | ||
| Line 109: | Line 109: | ||
The main applications of our system however, lie in fully exploiting its memorizing potential, as can be understood from the following: | The main applications of our system however, lie in fully exploiting its memorizing potential, as can be understood from the following: | ||
| - | + | ===Multipurpose Cell Lines=== | |
Our system can be trained to behave in a specific way, by setting its inducible toggle switch to one of its two states. This specific states can trigger specific and different events such as enzyme synthesis, transcriptional regulation, virion production, or even cell death. In this case, one can view the bacterial cell line containing this system, as a multipurpose cell line. One can add a certain chemical to a cell line, and train it to the desired behavior, instead of constructing two independent cell lines. | Our system can be trained to behave in a specific way, by setting its inducible toggle switch to one of its two states. This specific states can trigger specific and different events such as enzyme synthesis, transcriptional regulation, virion production, or even cell death. In this case, one can view the bacterial cell line containing this system, as a multipurpose cell line. One can add a certain chemical to a cell line, and train it to the desired behavior, instead of constructing two independent cell lines. | ||
| Line 117: | Line 117: | ||
For the purpose of creating a toggle that is activated in a specific phase, as is required for stable biological automatons, we introduced the concept of [https://2007.igem.org/ETHZ/Biology/parts#double_promoters double promoters] to the [http://partsregistry.org/Main_Page Registry of Standard Biological Parts], which can be helpful for future projects. As a concept, double promoters are expandable to handle multiple promoter sites, in the case of a greater number of toggles. | For the purpose of creating a toggle that is activated in a specific phase, as is required for stable biological automatons, we introduced the concept of [https://2007.igem.org/ETHZ/Biology/parts#double_promoters double promoters] to the [http://partsregistry.org/Main_Page Registry of Standard Biological Parts], which can be helpful for future projects. As a concept, double promoters are expandable to handle multiple promoter sites, in the case of a greater number of toggles. | ||
| - | =Link to Epigenetics= | + | ==Link to Epigenetics== |
Epigenetics refers to features like chromatin or DNA modifications that do not involve changes in the underlying DNA sequence and are stable over many cell divisions [1],[2]. If one has a closer look at our proposed system, one can also view it as a model-system for epigenetics: Although the DNA sequence itself stays the same, two different subpopulations of cells with different phenotypes can develop from it. Put simply, depending in which state (subpopulation) the toggle switch is, the cells will produce different fluorescent proteins upon addition of inducer molecules (aTc or IPTG). For example, if aTc is added one subpopulation will be red while the other will be yellow although both carry exactly the same DNA information. Therefore, the epigenetic feature here is the binding of specific repressor proteins whose production is dependent on the toggle switch state. | Epigenetics refers to features like chromatin or DNA modifications that do not involve changes in the underlying DNA sequence and are stable over many cell divisions [1],[2]. If one has a closer look at our proposed system, one can also view it as a model-system for epigenetics: Although the DNA sequence itself stays the same, two different subpopulations of cells with different phenotypes can develop from it. Put simply, depending in which state (subpopulation) the toggle switch is, the cells will produce different fluorescent proteins upon addition of inducer molecules (aTc or IPTG). For example, if aTc is added one subpopulation will be red while the other will be yellow although both carry exactly the same DNA information. Therefore, the epigenetic feature here is the binding of specific repressor proteins whose production is dependent on the toggle switch state. | ||
| - | =Team Members= | + | ==Team Members== |
[[Image:ETHZ_Group_photo_5.png|right|thumb|ETHZ iGEM2007 Team|300px]] | [[Image:ETHZ_Group_photo_5.png|right|thumb|ETHZ iGEM2007 Team|300px]] | ||
| Line 144: | Line 144: | ||
For more information about us, visit our [[ETHZ/Meet_the_team | Meet the Team]] page. | For more information about us, visit our [[ETHZ/Meet_the_team | Meet the Team]] page. | ||
| - | =Acknowledgments= | + | ==Acknowledgments== |
The idea for the project as well as its implementation was done by the ETH iGEM 2007 team. We would like to thank the people in [http://www.ipe.ethz.ch/laboratories/bpl/index Sven Panke's Lab], especially Andreas Meyer who was always there for us when we had a problem. Additionally, we would like to thank [http://www.facs.ethz.ch Alfredo Franco-Obregóns lab] and Oralea Büchi for the help with the flow cytometry. | The idea for the project as well as its implementation was done by the ETH iGEM 2007 team. We would like to thank the people in [http://www.ipe.ethz.ch/laboratories/bpl/index Sven Panke's Lab], especially Andreas Meyer who was always there for us when we had a problem. Additionally, we would like to thank [http://www.facs.ethz.ch Alfredo Franco-Obregóns lab] and Oralea Büchi for the help with the flow cytometry. | ||
| Line 159: | Line 159: | ||
</center> | </center> | ||
| - | =Site Map= | + | ==Site Map== |
In this wiki, we will present you a detailed description of the proposed system: starting with the [[ETHZ/Model | modeling of the system]], we describe both, [[ETHZ/Simulation | simulations and theoretical considerations]] of the system, as well as the actual [[ETHZ/Biology | implementation using bio-bricks]] accompanied by our [[ETHZ/Biology/Lab | lab notes]]. Additionally, you find some further [[ETHZ/Meet_the_team | information on the team]], some more details about [[ETHZ/Internal | ideas we developed]] before we came up with the system we finally implemented, and some [[ETHZ/Pictures | pictures]] documenting our work. | In this wiki, we will present you a detailed description of the proposed system: starting with the [[ETHZ/Model | modeling of the system]], we describe both, [[ETHZ/Simulation | simulations and theoretical considerations]] of the system, as well as the actual [[ETHZ/Biology | implementation using bio-bricks]] accompanied by our [[ETHZ/Biology/Lab | lab notes]]. Additionally, you find some further [[ETHZ/Meet_the_team | information on the team]], some more details about [[ETHZ/Internal | ideas we developed]] before we came up with the system we finally implemented, and some [[ETHZ/Pictures | pictures]] documenting our work. | ||
| Line 198: | Line 198: | ||
|} | |} | ||
| - | = References = | + | == References == |
[http://www.nature.com/nature/journal/v447/n7143/abs/nature05913.html;jsessionid=62903C604764B175945C03DB8639ECBD [1] Bird A] <i>"Perceptions of epigenetics"</i>, Nature 447:396-398, 2007 <br /> | [http://www.nature.com/nature/journal/v447/n7143/abs/nature05913.html;jsessionid=62903C604764B175945C03DB8639ECBD [1] Bird A] <i>"Perceptions of epigenetics"</i>, Nature 447:396-398, 2007 <br /> | ||
Revision as of 22:03, 25 October 2007
ETH Zurich - educatETH E.coli System