Valencia/Lab Work
From 2007.igem.org

|
|
Construction
We aimed for a straight-forward strategy that used all the parts from the registry of parts. From the computer work, we knew that in this system, time matters, so we focused on proteins that have a rapid degradation tag. This affects both repressors as well as the fluorescence proteins and this is the way how our system should work on a matter of minutes, not hours...
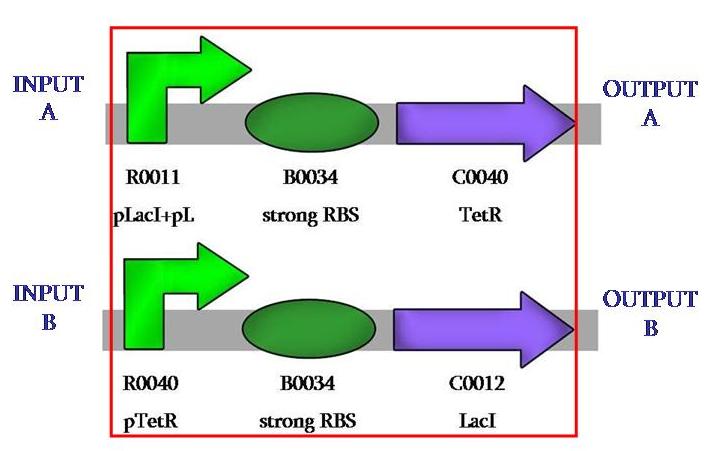
Our first achievement was to ligate the repressed promoter with the repressors, so to have two different constructions like this:
This is the comparator, a device that permits the modularity of the system, as one can use any desired input or ouput. In our case, we wanted to use fluorescence.
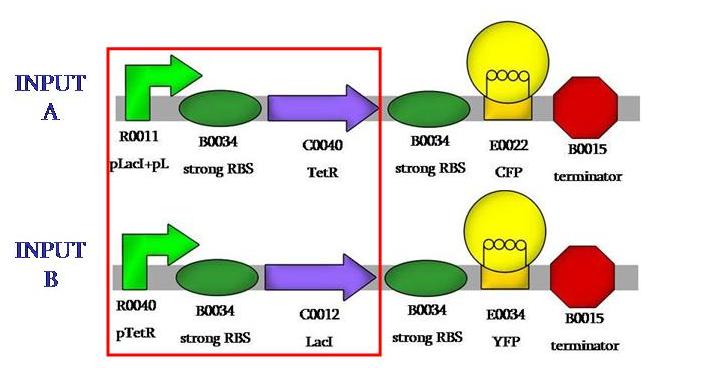
We aimed to assemble the fluorescence proteins backwards, those would be our output signals. We would have those constructs:
Now, we had to assemble the sensing part of our system, this would be the osmolarity-sensing promoters pOmpR and pOmpRm. we wanted to assemble them in order to build the E.coliRuler, the promoter calibrator device.
We first wantted to assemble pOmpR in both plasmids, so to validate the model and yield some system-specific parameters, then we wanted to compare pOmpR to pOmpRm.
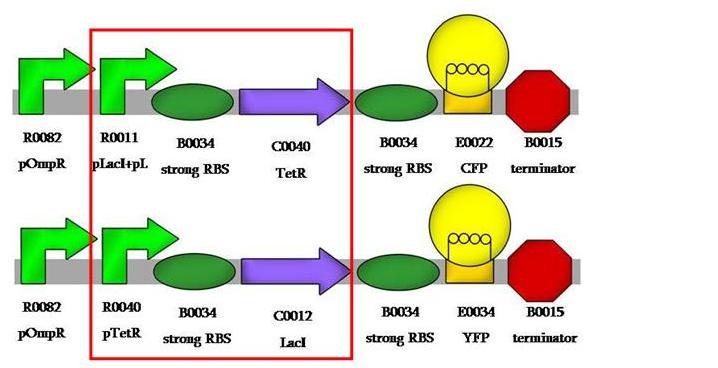
This would be our first system's design:
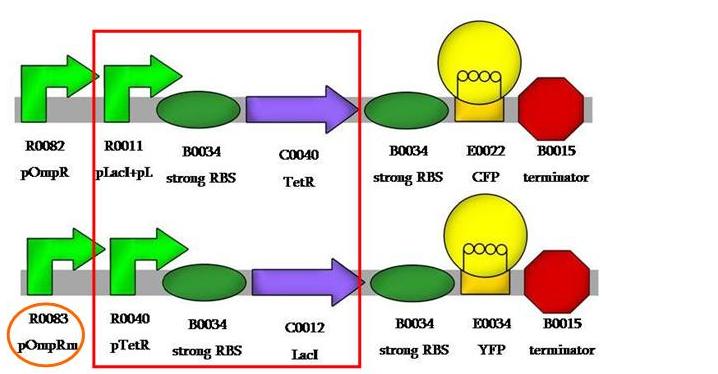
And, finally, we would like to have compared pOmpR to pOmpRm, like:
At last, we desired to have both constructs in one single bacterium, that is the reason why we had in mind to change the resistance backbone of one of the constructs to kanamicine. This way we would have a double selection for the finally transformed bacteria.
Fluorescence experiments
We planned to do two kinds of experiments.
First validating our model, using pOmpR as both inputs. And, then, compare pOmpR with pOmpRm.
You can take a closer look at it on our future experiments.