Valencia/Summary
From 2007.igem.org

|
|
While last year a vanillin sensor was designed (1), students from Universidad de Valencia and Universidad Politécnica de Valencia have gathered this year to create a device which behaviour resembles an electronic comparator.
The comparator works analyzing two input signals, like two proteins or metabolites, and responses with a given output, as fluorescence, related to the concentration of the input signal. Each input has an associated fluorescence colour.
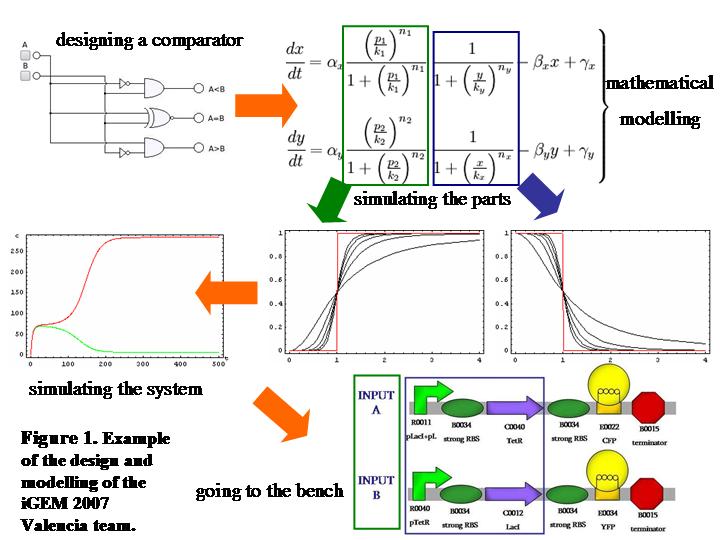
The interesting part is that differences of the output signals become amplified whenever a slight disparity between the initial concentrations of the input signals occurs. If inputs are equally sensed, both fluorescences have equal concentrations, but whenever one of the input’s levels is above the other, this fluorescence becomes greater and the other one gets inhibited. This behaviour emerges directly from the design of our system (Fig.1) in which we have used well established biological parts (2).
The potential modularity of the design was a positive aspect in order to develop this project. Inputs and outputs can be changed according to the goal of the specific project. For instance, changing the promoters this device could be used to compare whatever may be of interest for the scientist. Furthermore, the comparator device could be implemented in more complex systems, like a concentration stabilizer or a quality checker, verifying the right levels of an input. Additionally a system with two or more interconnected comparators could be designed to govern the production of a product of interest.
As a proof of principle, we have decided to calibrate promoters up to a standard. The focus was on a device that compares the promoter strengths of different mutants of the osmolarity-sensing promoter for the gene OmpC (3) and yields a fluorescent output with CFP and YFP. We have developed an effective model of the system to analyze in silico the behaviour of the system and study if it could be reliable.
- References:
1.- Rodrigo et al (2006). Vanillin cell sensor. IET Synth. Biol., 1, 74-78.
2.- Elowitz and Leibler (2000). A synthetic oscillatory network of transcriptional regulators. Nature, 403(6767), 335-8.
3.- Forst et al (1989). DNA-binding properties of the transcription activator (OmpR) for the upstream sequences of ompF in Escherichia coli are altered by envZ mutations and medium osmolarity. J. Bacteriol.,171 (6), 2949-2955.