Edinburgh/Yoghurt/Proof of concept
From 2007.igem.org
| Line 3: | Line 3: | ||
[[Edinburgh/Yoghurt| Introduction]] | [[Edinburgh/Yoghurt/Applications|Applications]] | [[Edinburgh/Yoghurt/Design|Design]] | [[Edinburgh/Yoghurt/Modelling|Modelling]] | [[Edinburgh/Yoghurt/Wet Lab|Wet Lab]] | [[Edinburgh/Yoghurt/Proof of concept|Proof of concept]] | [[Edinburgh/Yoghurt/References|References]] | [[Edinburgh/Yoghurt| Introduction]] | [[Edinburgh/Yoghurt/Applications|Applications]] | [[Edinburgh/Yoghurt/Design|Design]] | [[Edinburgh/Yoghurt/Modelling|Modelling]] | [[Edinburgh/Yoghurt/Wet Lab|Wet Lab]] | [[Edinburgh/Yoghurt/Proof of concept|Proof of concept]] | [[Edinburgh/Yoghurt/References|References]] | ||
---- | ---- | ||
| - | + | __TOC__ | |
In order to test the feasibility of gene expression in ''Lactobacillus'' and the possibility of making self flavouring yoghurt, we required a proof of concept. | In order to test the feasibility of gene expression in ''Lactobacillus'' and the possibility of making self flavouring yoghurt, we required a proof of concept. | ||
Revision as of 17:53, 26 October 2007
 Introduction | Applications | Design | Modelling | Wet Lab | Proof of concept | References
Introduction | Applications | Design | Modelling | Wet Lab | Proof of concept | References
Contents |
In order to test the feasibility of gene expression in Lactobacillus and the possibility of making self flavouring yoghurt, we required a proof of concept.
As both the pigment and flavour pathways are rather complex and will require modification and optimisation before their expression in yoghurt we planned to use the much simpler Red Fluorescent Protein (RFP) gene as a proof of concept.
So far we have managed to transform a number of RFP-pTG262 vectors containing a variety of promoters into E. coli and also Bacillus subtillis, a gram positive bacterium.
We chose to initially transform Bacillus subtillis with our proof of concept vectors over Lactobacillus for two reasons:
1. the bacterium is naturally competent at certain stages of its life cycle and therefore has a much simpler transformation process
2. members of the French lab have had successful results from when they previously worked with Bacillus subtilis
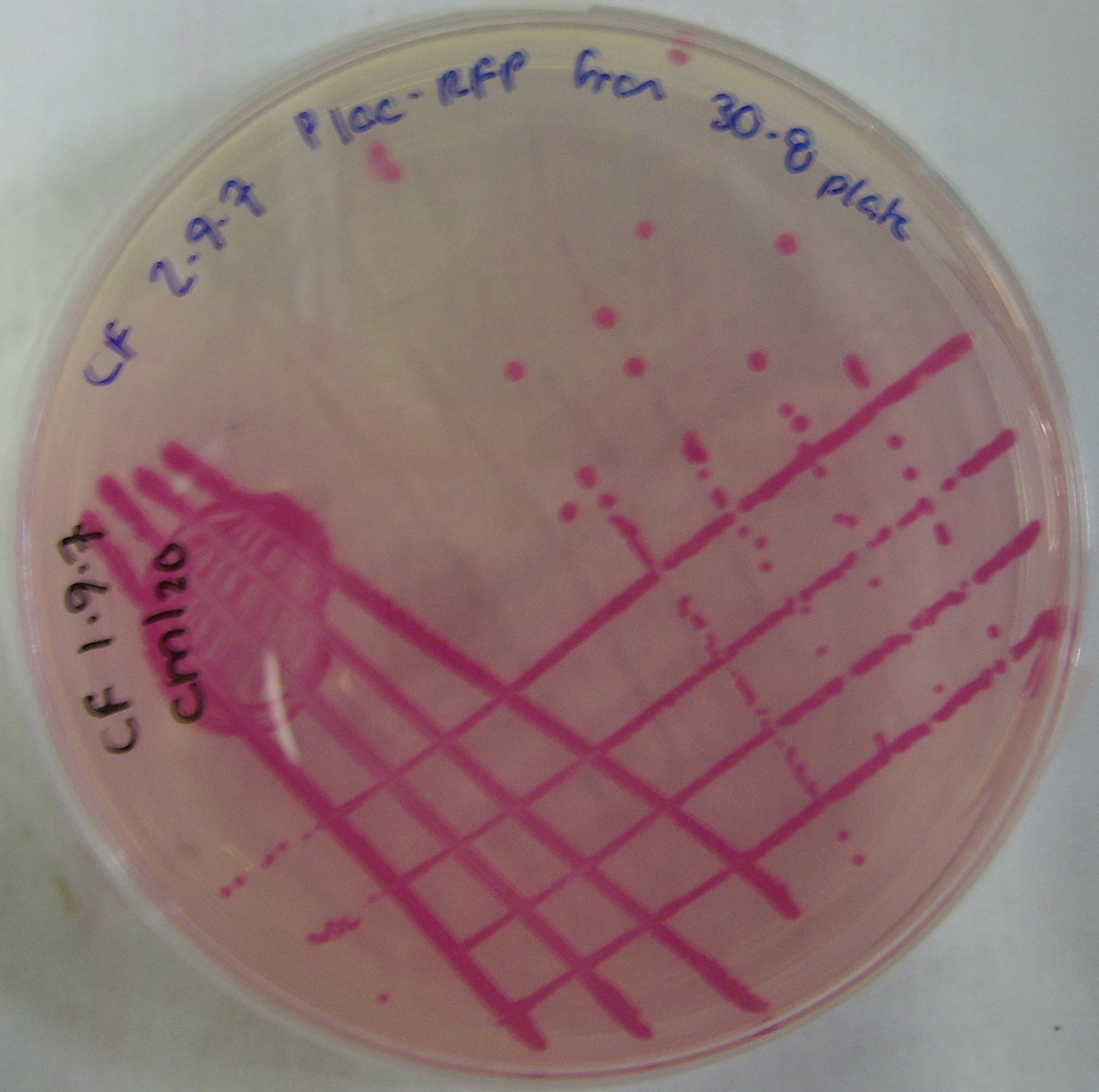
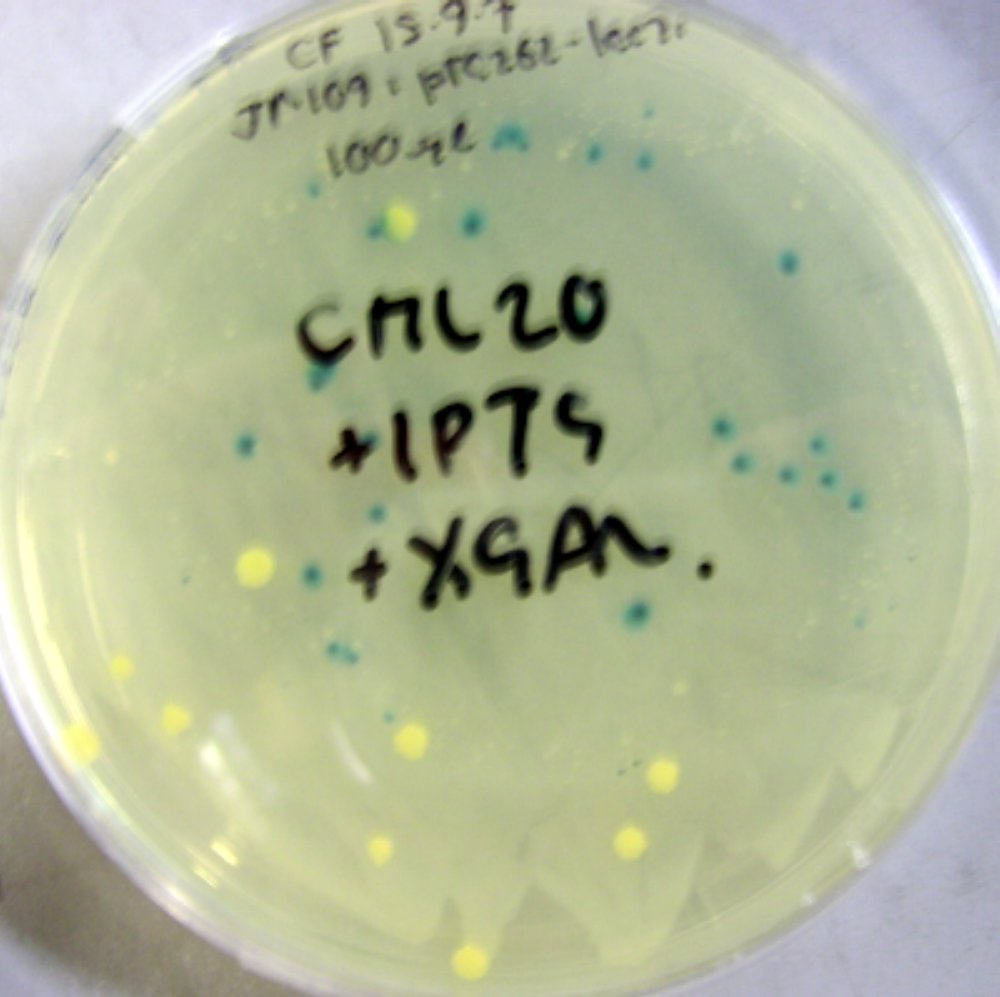
pTG262-PLac-RFP construct
We have inserted the lactose induced RFP gene into the pTG262 vector
- this vector was then transformed into E. coli
- growth on IPTG-Xgal media resulted in RFP expression and production of red colonies, these can be viewed in figure 1
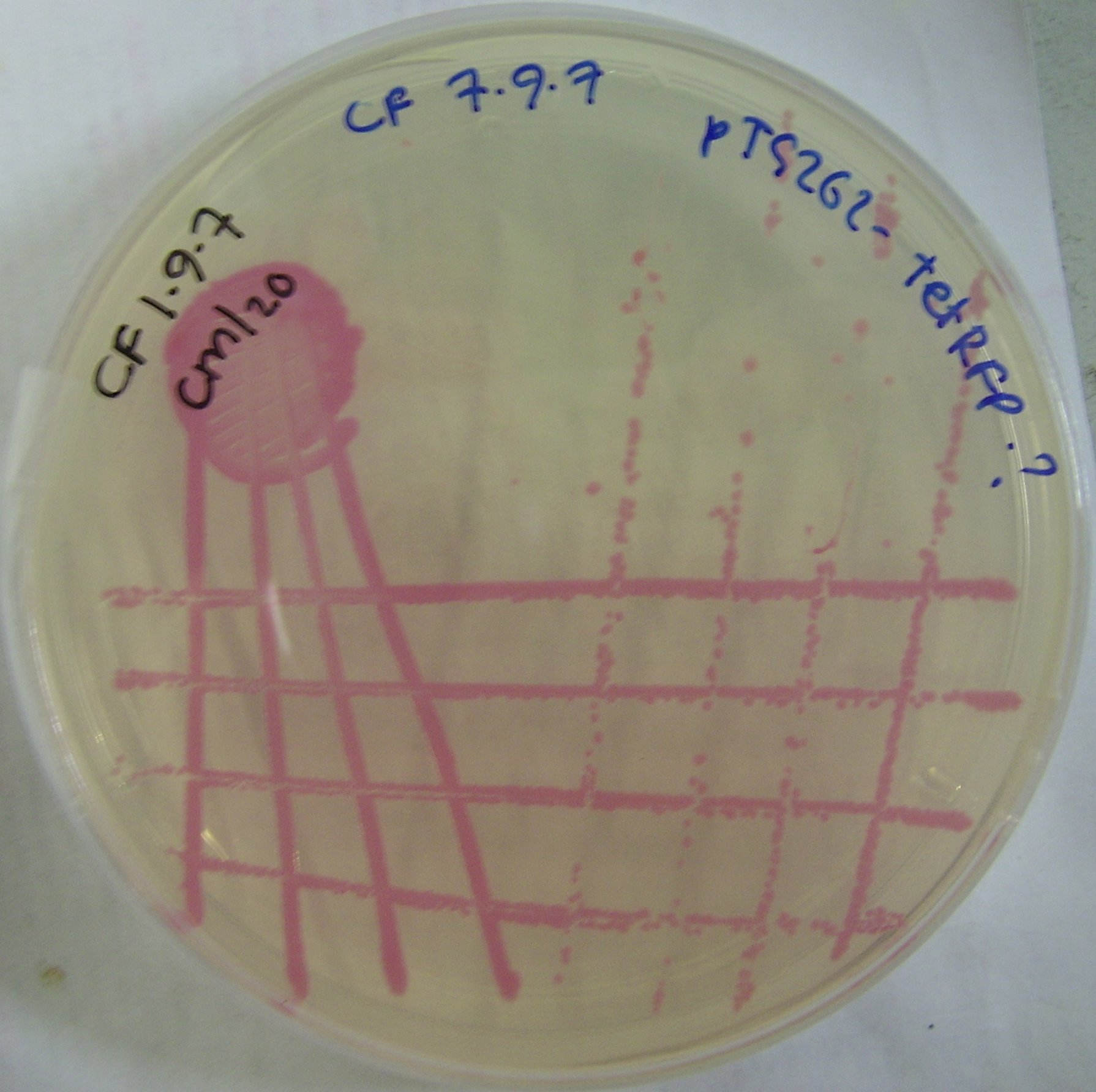
pTG262-Ptet-RFP construct
We ligated the Ptet induced RFP biobrick into the pTG262 vector. We then transformed the vector into E. coli.
RFP synthesis was observed.
ptet incuded RFP E. coli colonies can be viewed in figure 2.
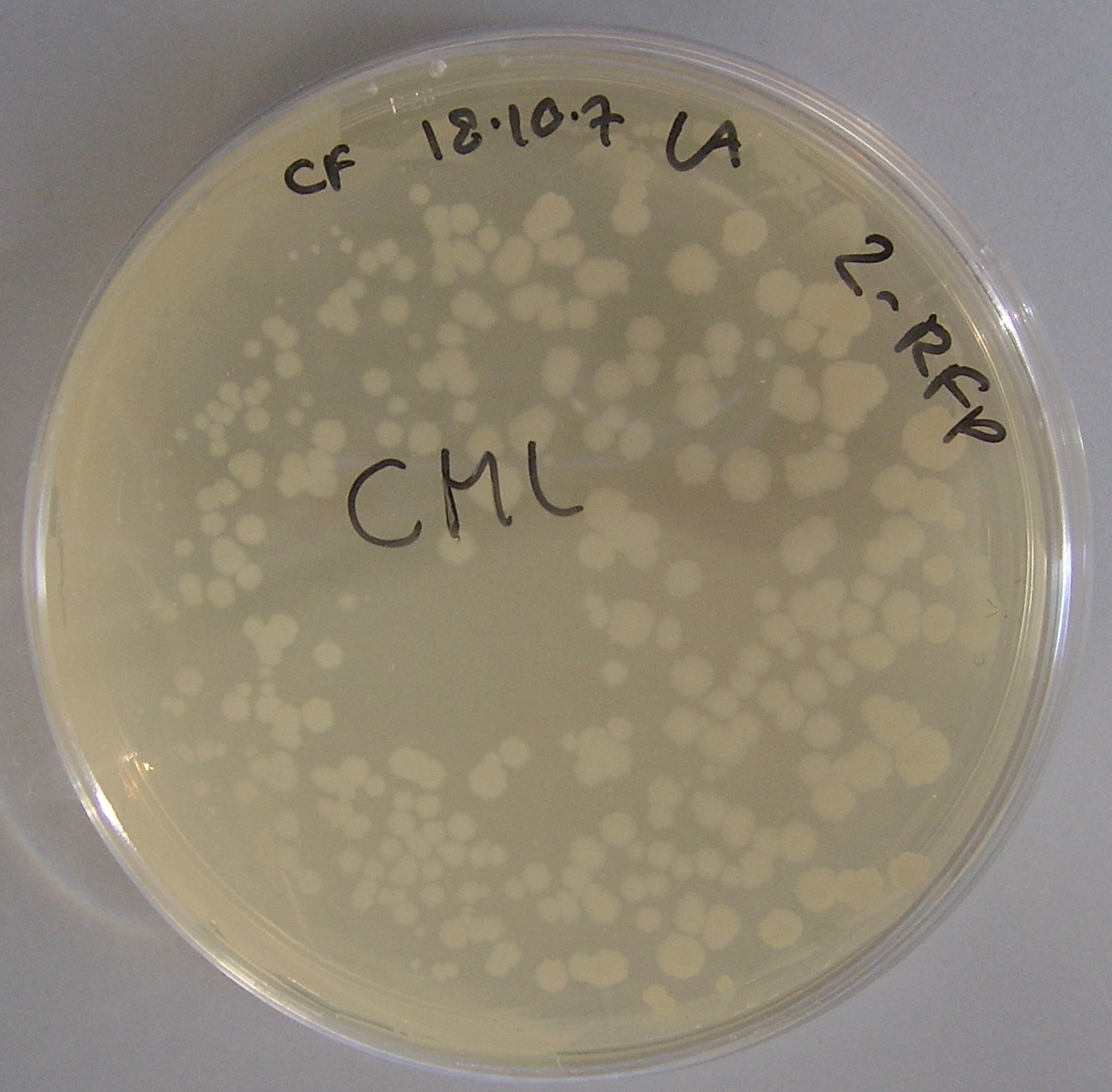
pTG262 expression in Bacillus subtilis
After the successful expression of both the pTG262 RFP constructs in E. coli, we decided to determine if pTG262 could be stably maintained in Bacillus subtilis.
To do this we transformed both the pTG262 and Plac-RFP-pTG262 vectors into Bacillus subtilis.
Figure 3 depicts the successful transformation and expression of pTG262 in Bacillus subtilis.
This is determined by the ability of Bacillus to grow on chloramphenicol plates.
Unfortunately RFP was not expressed from the pLac promoter, as shown by the colonies being white not red. We are attributing this to the Bacillus transcription machinary (particularly RNA polymerase and its associated sigma factors) not recognising the promoter.
Introduction | Applications | Design | Modelling | Wet Lab | Proof of concept | References