Melbourne/Lab Notebook gv 2
From 2007.igem.org
< Melbourne(Difference between revisions)
| Line 1: | Line 1: | ||
| + | [[Melbourne/Lab GV Notebook| <Return to lab book summary>]] | ||
==Prepared for site dirrected mutagenesis== | ==Prepared for site dirrected mutagenesis== | ||
*resuspended primers supplied by geneworks to produce stock solution at 125ng/uL using filtered,autoclaved,milliQ water. | *resuspended primers supplied by geneworks to produce stock solution at 125ng/uL using filtered,autoclaved,milliQ water. | ||
Latest revision as of 12:31, 26 October 2007
Prepared for site dirrected mutagenesis
- resuspended primers supplied by geneworks to produce stock solution at 125ng/uL using filtered,autoclaved,milliQ water.
- Produced working solutions at 125ng/2uL by diluting 10uL of stock with 190uL of filtered,autoclaved,milliQ water.
- Produced NZY broth 100mL autoclaved
- Produced 1mL of 10mM IPTG filter sterilised
- Produced 1mL of 2% XGAL in DMF(Dimethylformamide) filter sterilized
Site dirrected Mutagenesis Round #1
- Applied the stratagene Site directed mutagenesis protocolto the following DNA and primer pairs, which when plated out on LB AMP plates producing the numbers of colonies shown. Several of these were picked and tubes marked with a character suffix as shown in table.
| Mutation Number | Template DNA (10ng) | Sence Primer | Antisence Primer | #Colonies | Picks named |
|---|---|---|---|---|---|
| 1 | pWhitescript | control #1 | control #2 | Blue and White on XGAL&IPTG plates | |
| 2 | pNL29T1 | GvpL-g213a | GvpL-g213a-R | None | |
| 3 | pNL29T1 | GvpL-g318a | GvpL-g318a-R | 3 | 3A,3B,3C |
| 4 | pNL29T1 | JMY-g318a | JMY-g318a-R | 26 | 4A,4B,4C |
| 5 | pNL29T1 | Comp-g318a | Comp-g318a-R | 60 | 5A,5B,5C |
| 6 | pNL29T1 | GvpL-g351a | GvpL-g351a-R | 1 | 6A |
| 7 | pNL29T1 | GvpL-g696a | GvpL-g696a-R | None | |
| 8 | pNL29T1 | GvpA-t57c | GvpA-t57c-R | None | |
| 9 | pNL29T1 | GvpQ-g150a | GvpL-Q-g150a | None | |
| 10 | pNL29T1 | GvpQ-g183a | GvpQ-g183aa-R | 6 | 10A,10B,10C,10D,10E |
| 11 | pNL29T1 | GvpP-g441a | GvpP-g441a-R | 200 | 11A,11B,11C,11D |
| 12 | PUC18 | N/A | N/A | 30 Blue on Xgal&IPTG Plates | |
| 13 | N/A | N/A | N/A | None | |
| 14 | pNL29T1 | GvpL-g213a | GvpL-g213a-R | None | |
| 15 | IPTG & Xgal | N/A | N/A | None |
- Picked colonies from Plates and culture overnight
- Produced glycerol stocks from 900uL of each overnight culture.
- Minipreped remaining culture (4mL) giving 100uL.
- Measure DNA concentrations using 2uL of miniprep(100uL) diluted in 98uL of TE.
| History | Mutation tube\colony: | A | B | C | D | E |
|---|---|---|---|---|---|---|
| pNL29T1-(GvpP-g441a)-> | 3 | 141 | 112 | |||
| pNL29T1-(GvpL-g318a)-> | 4 | 108 | 218 | |||
| pNL29T1-(Comp-g318a)-> | 5 | 141 | 119 | |||
| pNL29T1-(GvpL-g351a)-> | 6 | 212 | ||||
| pNL26P3-(GvpQ-g183a)-> | 10 | 305 | 253 | 258 | 392 | 272 |
| pNL26P3-(GvpP-g441a)-> | 11 | 221 | 190 | 259 | 204 |
- Diagnostic digests were performed using
- A) 21.5uL of each in Buffer3 with 1uL PstI (20U) at 37degC for 2h30min. (25uL reaction)
- B) 21.5uL of each p29T1 based colony in Buffer2 with 1uL EcoRI (20U) and 1uL BamHI (20U) at 37degC for 2h30min. (26uL reaction)
- C) 21.5uL of each pNL26P3 based colony and in Buffer2 with 1uL EcoRI (20U) at 37degC for 2h30min. (25uL reaction)
- Prepared two 20 lane 100mL agarose gel
- Loaded 20uL of digest
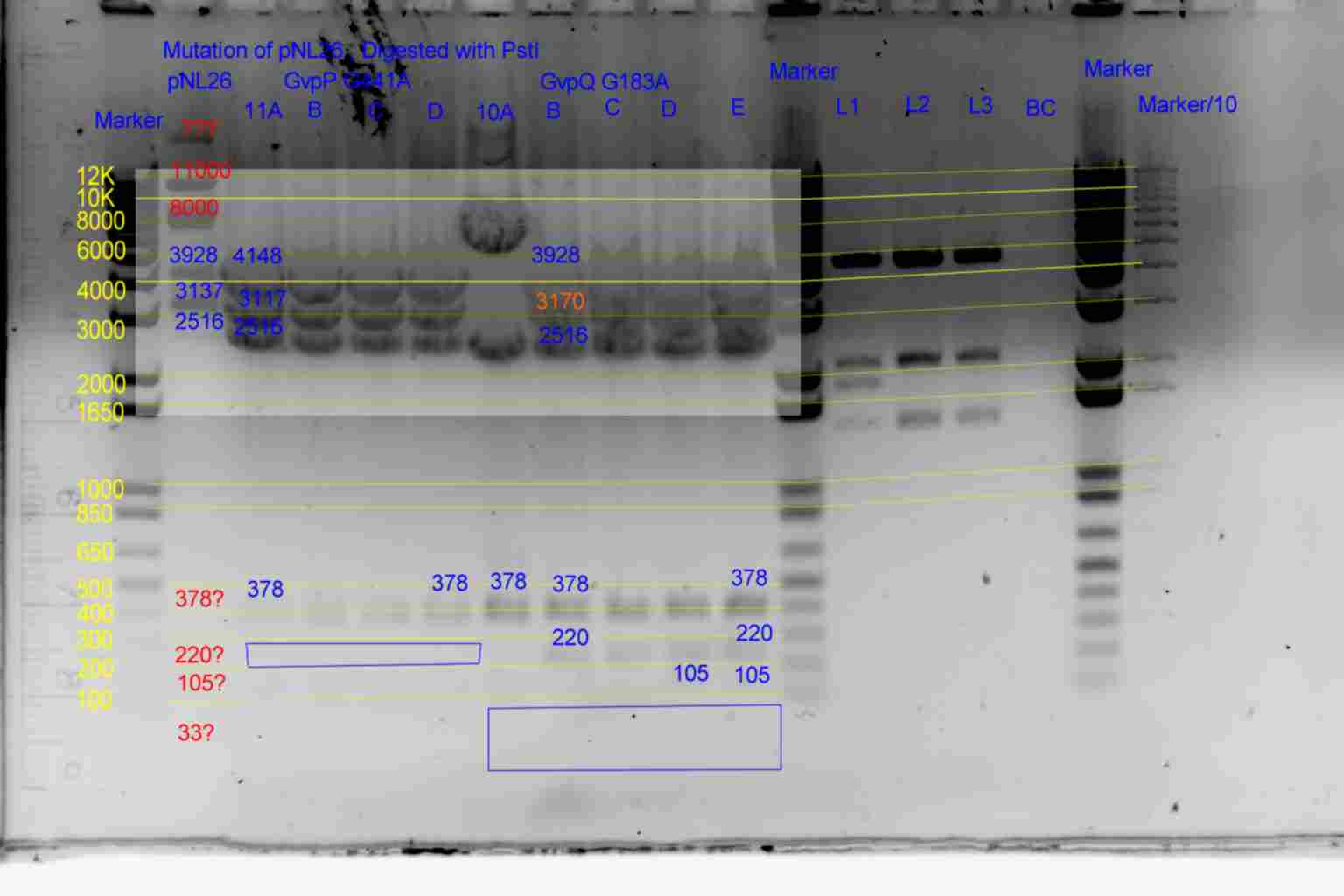
- Digest Pattern
- Expected effects
| History | Mutation tube | FRAGMENTS | |||||||
|---|---|---|---|---|---|---|---|---|---|
| pNL29T1 | pNL29T1 | 5983 | 2516 | 378 | 105 | ||||
| pNL29T1-(GvpP-g441a)-> | 3 | 6088 | 2516 | 378 | <- | ||||
| pNL29T1-(GvpL-g318a)-> | 4 | 5983 | 2516 | 483 | <- | ||||
| pNL29T1-(Comp-g318a)-> | 5 | 5983 | 2516 | 483 | <- | ||||
| pNL29T1-(GvpL-g351a)-> | 6 | 5983 | 2516 | 378 | 105 | ||||
| pNL26P3 | pNL26P3 | 3928 | 3137 | 2516 | 378 | 220 | 105 | 33 | |
| p26P3-(GvpQ-g183a)-> | 10 | 3928 | 3170 | 2516 | 378 | 220 | 105 | <- | |
| p26P3-(GvpP-g441a)-> | 11 | 4148 | 3137 | 2516 | 378 | <- | 105 | 33 |
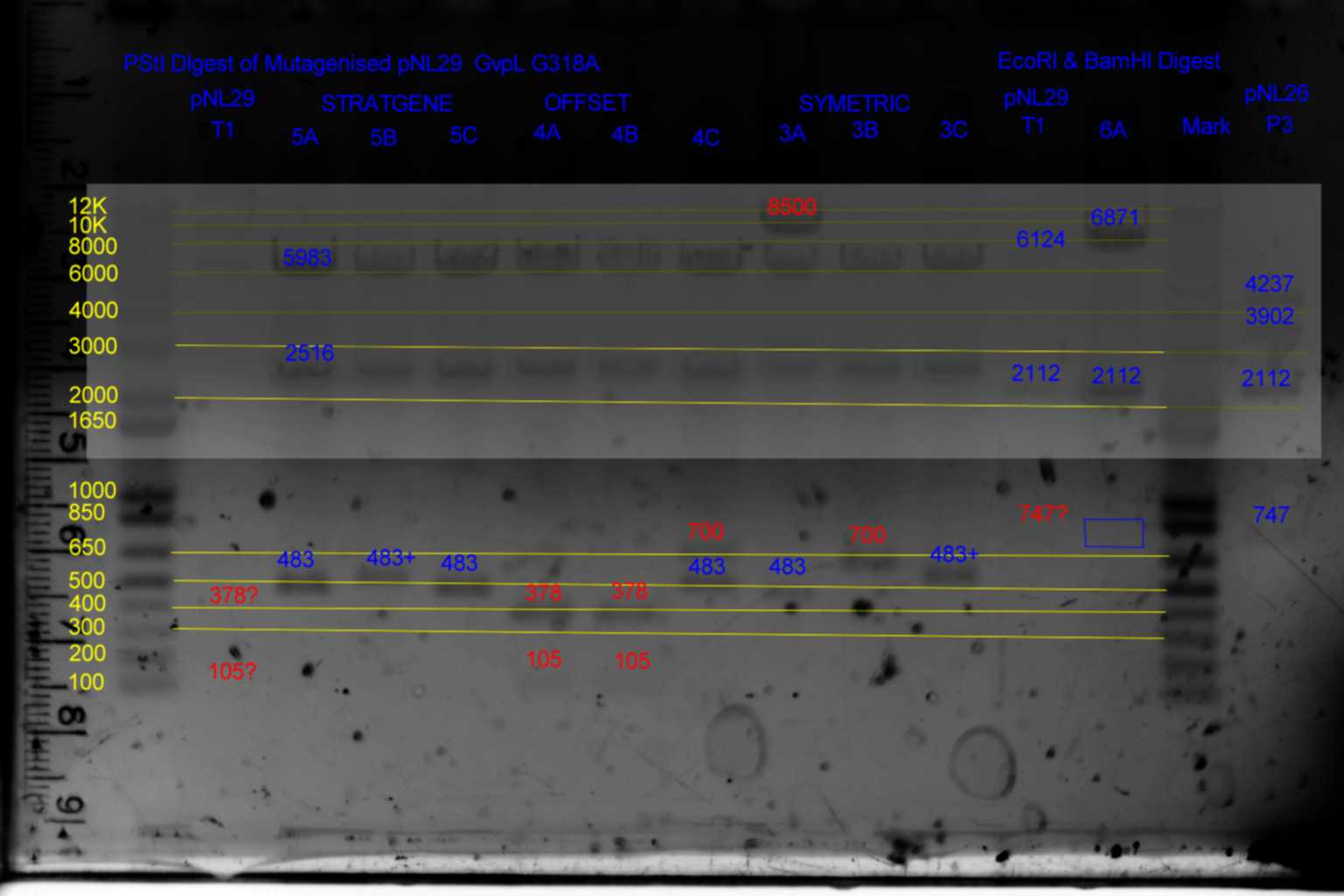
| History | Mutation tube | Fragments: | |||
|---|---|---|---|---|---|
| pNL29T1 | pNL29T1 | 6124 | 2112 | 747 | |
| pNL29T1-(GvpP-g441a)-> | 3 | 6124 | 2112 | 747 | |
| pNL29T1-(GvpL-g318a)-> | 4 | 6124 | 2112 | 747 | |
| pNL29T1-(Comp-g318a)-> | 5 | 6124 | 2112 | 747 | |
| pNL29T1-(GvpL-g351a)-> | 6 | 6871 | 2112 | <- |
- Results of PstI diagnostics:
- pNL29 control was barely visable.
- Bands less than 300bp were not visable
- Colonies 5A,5B,5C,and 3C all showed bands at 483bp and not at 378bp as required although 5B and 3C appear slightly larger than the other two.
- Colony 6A shows no band at 747bp and a band at 6871bp as required.
- Colonies 4A,and 4B contain a band at 378bp indicating failure of mutagenesis.
- The following colonies show additional bands of unknown origin: 3B and 4C (700bp) , 3A (8500bp) of unknown origin.
- See Gel
- Conclude that 5A and 6A are suitable templates for the second round of mutagenesis.
- Results of EcoRI diagnostics:
- pNL26 control barely visable.
- Colonies 11A,B,C,D and 10A,B,C,D,E all show required pattern but the 33bp band is unlikely to be visible making the 10A..E test inconclusive, and the 220bp band in 11A..D is only barely visible in 11A..E making its diagnosis also risky.
- See Gel
- Conclude that 10B and 11A may be suitable templates for the second round of mutagenesis.
- Conclude that Primers generated by the stratagene program Comp-g318a outperformed (60 colonies of which 2/3 provided correct product ) both offset primers JMY-g318a (26 colonies of which 2/3 gave template DNA)primers and hand designed primers Gvp-g318a (3 colonies of which when using the stratagene PCR protocol.
| Primers name | Design protocol | no colonies | #correctly mutated | #unknown | # template |
|---|---|---|---|---|---|
| (Comp-g318a) | Statagene | 60 | 2 | 1 | - |
| (JMY-g318a) | Offset | 26 | - | 1 | 2 |
| (GvpL-g318a) | Hand design | 3 | - | 3 | - |
- It was however also noted that the success of the PCR with the hand generated primers was negatively correlated with the melting temperature of the primers.
- It was therefore decided to increase the initial melting time from 1 to 2 minutes and increase subsequent melting times from 50 seconds to 1 minute in the next round of mutagenesis.