Imperial/Infector Detector/F2620 Comparison
From 2007.igem.org
m (→Summary of Comparison) |
m (→Summary of Comparison) |
||
| Line 47: | Line 47: | ||
|<center>[[image:In vitro in vivo comp.png|700px]]</center> | |<center>[[image:In vitro in vivo comp.png|700px]]</center> | ||
|- | |- | ||
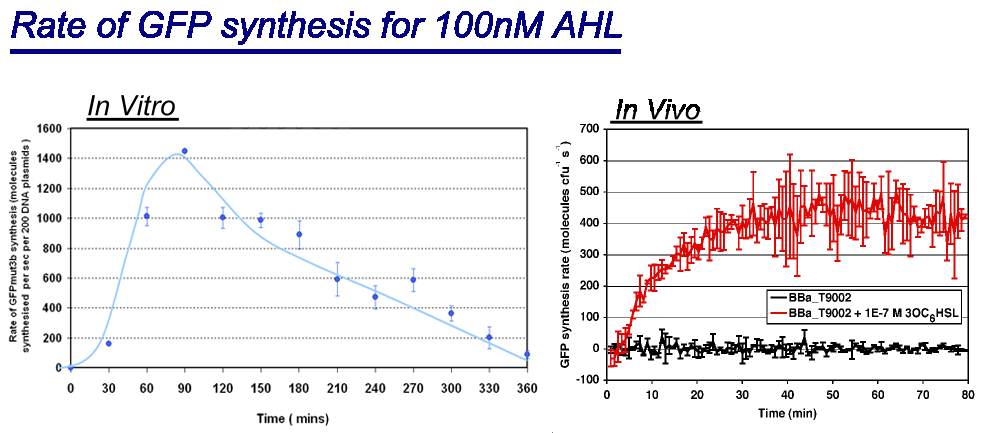
| - | |style="text-align: left;" |Comparison between ''in vivo'' and ''in vitro'' for rate of GFPmut3b synthesis for 100nM AHL. The ''in vivo'' chassis used was the bacterial strain MG1655 and the ''in vitro'' chassis was Promega Commercial S30 Cell Extract | + | |style="text-align: left;" |Comparison between ''in vivo'' and ''in vitro'' for rate of GFPmut3b synthesis for 100nM AHL. The ''in vivo'' chassis used was the bacterial strain MG1655 and the ''in vitro'' chassis was Promega Commercial S30 Cell Extract(60µl) |
<br> | <br> | ||
The graph shows the following: | The graph shows the following: | ||
| - | *''in vivo'' has a maximal rate of 400-500 molecules of GFP synthesised per second per cell. In addition the rate reaches a steady state after around 30minutes and maintains it for the duration of the testing.<br> | + | *''in vivo'' has a '''maximal rate of 400-500 molecules of GFP synthesised per second per cell'''. In addition the rate reaches a steady state after around 30minutes and maintains it for the duration of the testing.<br> |
| - | *''in vitro'' has the equivalent of | + | *''in vitro'' has the equivalent of 1400 molecules of GFP synthesised per second per cell equivalent, the cell equilavent being based upon the normalization of DNA plasmids. Interestingly the ''in vitro'' chassis does not reach a steady state, in fact it decreases in rate of synthesis after 90 minutes and keeps decreasing until rate is zero at around 360 minutes. |
<br> | <br> | ||
The reason why the ''in vitro'' chassis never reaches a steady state is because of the limited energy and metabolites available, this is unlike ''in vivo'' which is supported by the media that it is grown upon. | The reason why the ''in vitro'' chassis never reaches a steady state is because of the limited energy and metabolites available, this is unlike ''in vivo'' which is supported by the media that it is grown upon. | ||
Revision as of 21:21, 26 October 2007

Summary of Comparison
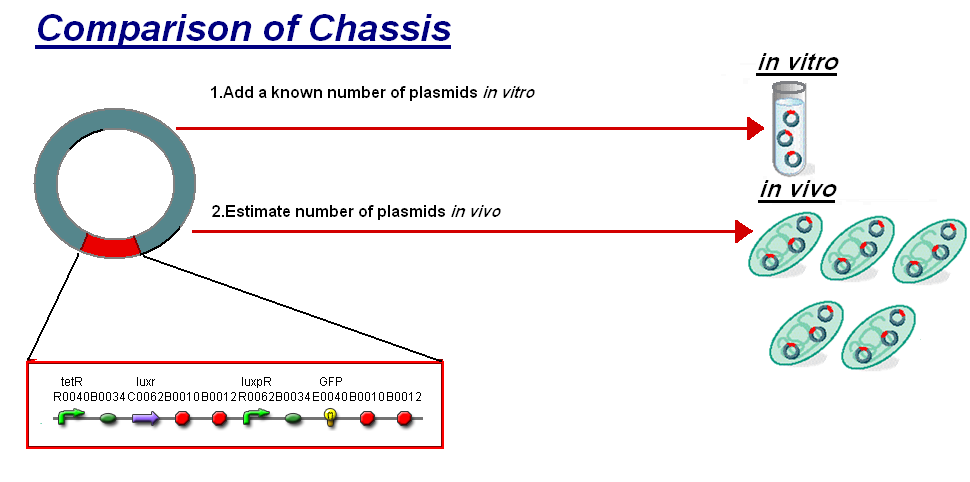
We thought to compare our in vitro characterisation to the characterisation of F2620 in vivo with the aim to highlight some of the differences between the chassis and investigate how the constructs characteristics may change between them. The F2620 is an ideal construct to compare for comparison because of its detailed characterisation in vivo. The construct is the same as the construct 1 that was used for infecter detector, pTet-LuxR-pLux-GFPmut3b. The key results the comparison were;
- The creation of a new unit to allow comparison between in vitro and in vivo chassis.
- That although we are changing the E.coli chassis from in vivo to in vitro the construct characteristic response is independent of the chassis.
These are exciting findings, revealing the potential for the exploration of new chassis and the ability to use constructs in an exchangable mannor.
|
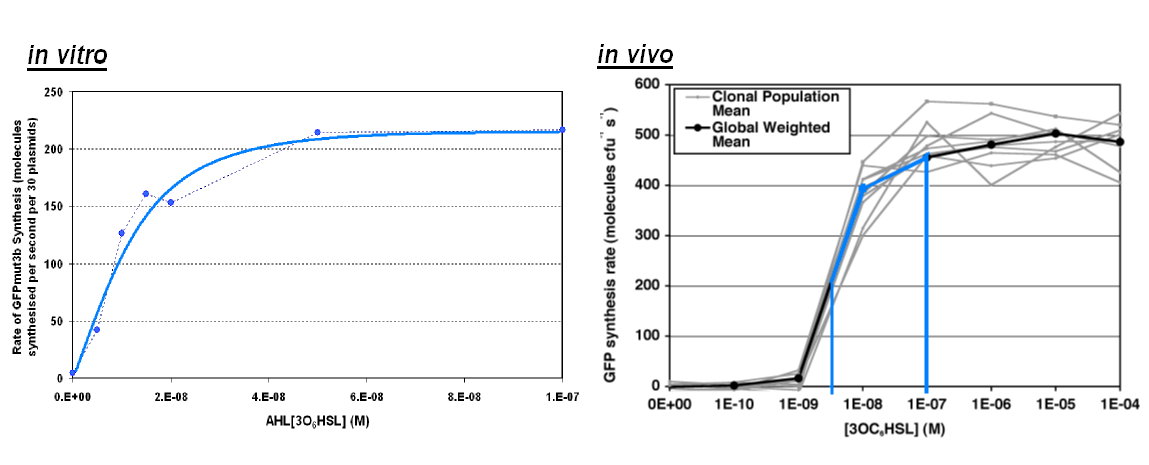
Transfer Function