Imperial/Infector Detector/Testing
From 2007.igem.org
m (→Aims) |
m |
||
| Line 22: | Line 22: | ||
== Aims of Testing== | == Aims of Testing== | ||
| - | To test and characterise key characteristics of our system e.g. the sensitivity of the system to AHL. To do this, we induce the system with known concentrations of AHL and measure the fluorescence output. | + | To test and characterise key characteristics of our system e.g. the sensitivity of the system to AHL. To do this, we induce the system with known concentrations of AHL and measure the fluorescence output. Then using a calibration curve the fluorescence was converted into the number of GFPmut3b molecules synthesised, click on the following link for an explaination about how to use the calibration curve. The full results and protocols can be found on the links [[Imperial/2007/Wet_Lab/results/ID3.1|results]] and [[Imperial/Wet_Lab/Protocols/ID3.1|protocol]] pages. |
| - | + | ||
| - | + | ||
| - | + | ||
=Results= | =Results= | ||
| Line 31: | Line 28: | ||
|<center><br>[[Image:GFPMolecule syn ID2 Final.PNG|500px]]</center> | |<center><br>[[Image:GFPMolecule syn ID2 Final.PNG|500px]]</center> | ||
|- | |- | ||
| - | | | + | |The results show us the following: |
| - | + | ||
| - | The results show us the following: | + | |
*The output of '''GFPmut3b increases with input of AHL''' | *The output of '''GFPmut3b increases with input of AHL''' | ||
*The system is sensitive to a range of '''5-1000nM AHL''' | *The system is sensitive to a range of '''5-1000nM AHL''' | ||
Revision as of 10:39, 24 October 2007

Infector Detector: Testing
Aims of Testing
To test and characterise key characteristics of our system e.g. the sensitivity of the system to AHL. To do this, we induce the system with known concentrations of AHL and measure the fluorescence output. Then using a calibration curve the fluorescence was converted into the number of GFPmut3b molecules synthesised, click on the following link for an explaination about how to use the calibration curve. The full results and protocols can be found on the links results and protocol pages.
Results
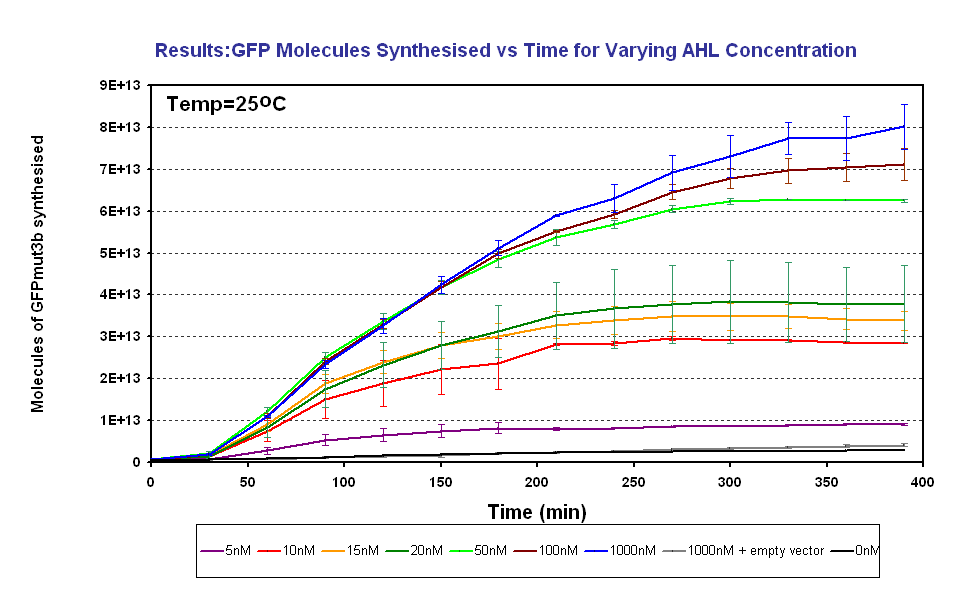 |
The results show us the following:
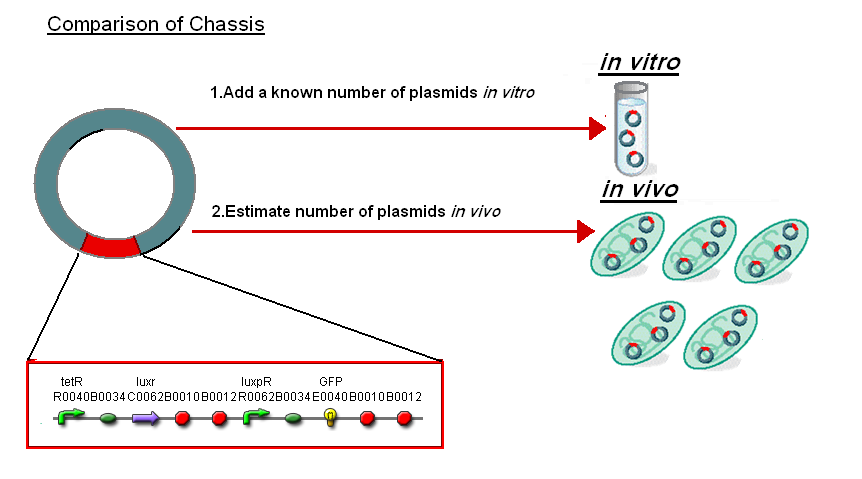
In addition the results in vitro have been compared to the work on BBa_F2620(pTet-LuxR-pLux-GFPmut3b in vivo which is the same as the construct 1 used for infecter detector. To do this we investigated the comparison between in vitro and in vivo. |
Normalise the in vitro on the plasmids to give a platform for comparison:
- In Vitro - 4µg of DNA was added which for pTet-LuxR-pLux-GFPmut3b is 904823007 plasmids
- In Vivo - Each cell has ~30 plasmids per cell
To compare we normalised the data of in vitro GFPmut3b molecules synthesised per 30 plasmids to allow some comparison to the in vivo data.
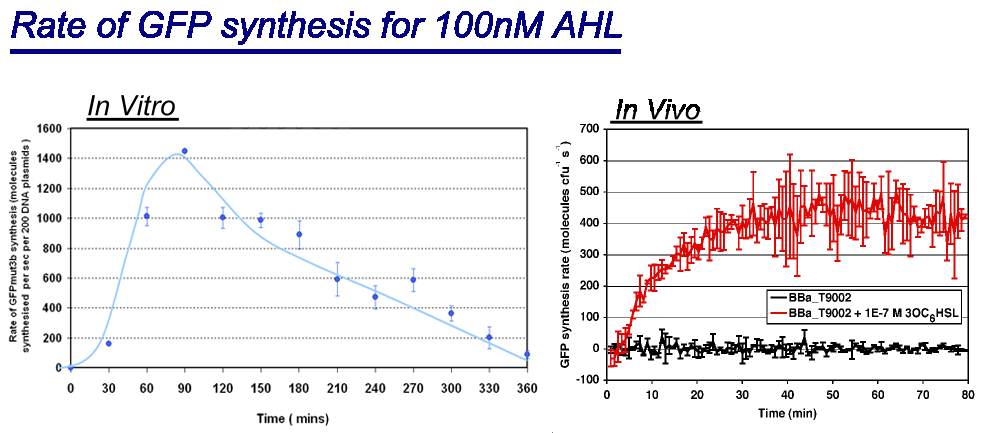
Rate of GFPmut3b Synthesis for 100nM AHL
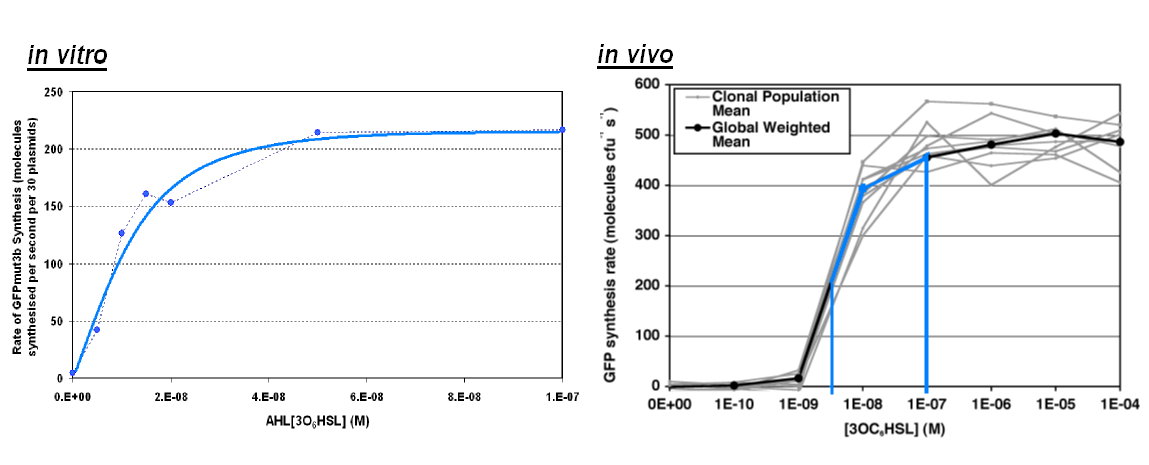
Transfer Function
Summary
Below is list of which of the orginial Specifications that our infecter detector achieved:
| Achievements | ||
| Inputs | Sensitive to 5-1000nM | |
| Outputs | Future work - Using Stronger fluorescent protein such as DsRed express | |
| Response Time | Systems responds <30minutes | |
| Operating Conditions | System works at 25°C | |
| Health & Safety | Cell Free in vitro chassis | |
| Lifespan | Can be stored in freezer for prolonged periods | |
| Packaging | Future Work - Using our chassis in a spray |