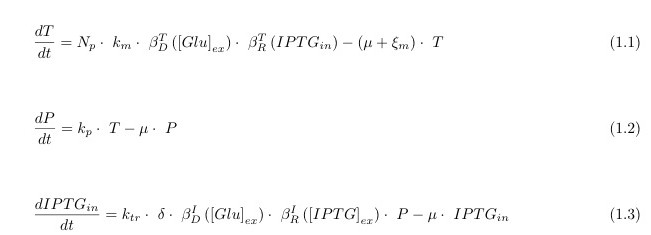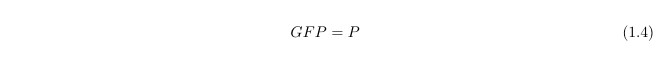Bologna
From 2007.igem.org
(→Introduction) |
Guido Costa (Talk | contribs) (→Experimental SetUp) |
||
| Line 147: | Line 147: | ||
The control software is implemented in a Labview environment and permits the regulation of the excitation wavelength and the calibration of the system. It also pictures the output signal from the photomultiplier, which can be memorized and elaborated. | The control software is implemented in a Labview environment and permits the regulation of the excitation wavelength and the calibration of the system. It also pictures the output signal from the photomultiplier, which can be memorized and elaborated. | ||
| + | |||
| + | |||
| + | ===Image Acquisition=== | ||
| + | |||
| + | Using the experimental set up illustrated [https://2007.igem.org/Bologna_University/Microscopy:_Experimental_Set_Up here], we obtain short movies of fluorescent bacteria; then we extract several frames from each of them, using a dedicated software. Example images are shown below. | ||
| + | |||
| + | |||
| + | [[Image:orig100.jpg|left]] | ||
| + | |||
| + | [[Image:orig200.jpg|absright]] | ||
| + | |||
| + | |||
| + | |||
| + | ===Image Elaboration=== | ||
| + | |||
| + | Since we need to know how much of the total image area is occupied by bacteria, we process the images with our segmentation algorithm implemented in Matlab. | ||
| + | |||
| + | Here are the processing phases: | ||
| + | |||
| + | *the RGB image is read and the green channel is extracted; | ||
| + | *a morphological top hat filtering is performed on the grayscale image; | ||
| + | *by means of an adaptive threshold, the image is tranformed in a binary one; | ||
| + | *the distance between every point and the nearest black pixel is calculated; | ||
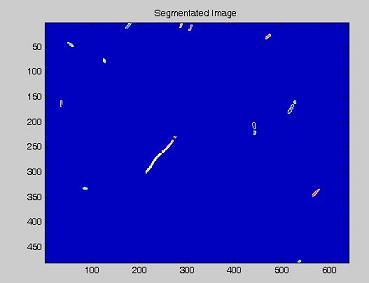
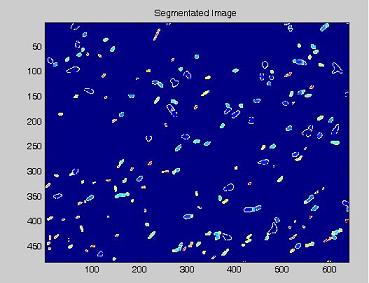
| + | *on this last image the [http://en.wikipedia.org/wiki/Watershed_%28algorithm%29 watershed] algorithm is applied: every pixel is assigned a label, depending on the segmentated region it belongs; then, every labeled region is represented with a different colour, as in the images shown below; | ||
| + | |||
| + | |||
| + | [[Image:segm100.jpg|left]] | ||
| + | |||
| + | [[Image:segm200.jpg|absright]] | ||
| + | |||
| + | |||
| + | *the area of every segmented region is calculated, checking that the summation of these areas (including the background) balances the image total area; | ||
| + | *two complementary matrices are created: | ||
| + | -''ImageWithOutBackground'', containing the intensity positive values corresponding only to pixels recognised as bacteria, with zeros elsewhere; | ||
| + | -''ImageBackground'', which contains the intensity positive values of pixels recognised as background, with zeros as other entries; | ||
| + | *with a summation over all the ''ImageWithOutBackground'' matrix' entries, the total intensity is obtained; | ||
| + | *dividing this value by the total bacteria area, the output is the normalized intensity we use to compare the fluorescence of different kinds of bacteria. | ||
| + | |||
| + | ''Thanks to Camilo Melani for his competence and kindness during the algorithm implementation.'' | ||
==Biodevice== | ==Biodevice== | ||
Revision as of 11:15, 26 October 2007
"Ὁ βίος βραχὺς, ἡ δὲ τέχνη μακρὴ, ὁ δὲ καιρὸς ὀξὺς, ἡ δὲ πεῖρα σφαλερὴ, ἡ δὲ κρίσις χαλεπή"
"Vita brevis, ars longa, occasio praeceps, experimentum periculosum, iudicium difficile"
"Art is long, life is short, opportunity is fleeting, experience is deceitful, judgement is difficult"
"La vita è breve, l'arte è lunga, l'occasione fuggevole, l'esperimento pericoloso, il giudizio difficile"
(Aforisma di Ippocrate di Coo)
Contents |
About Us
Welcome to Bologna’s IGEM Wiki!
Our team consists of Francesco Pasqualini, Guido Costa, Iros Barozzi, Silvia Tamarri, Francesca Buganè, and Michela Mirri, all soon-to-be juniors majoring in Biomedical Engineering at Bologna University; and Alice Pasini, a just graduated student.
We are advised by [http://www.ing2.unibo.it/Ingegneria+Cesena/Strumenti+del+Portale/Cerca/paginaWebDocente.htm?NRMODE=Published&TabControl1=TabContatti&UPN=emanuele.giordano%40unibo.it Dr Emanuele Giordano], Lecturer in Biochemistry and [http://www-micro.deis.unibo.it/cgi-bin/user?tartagni Prof. Marco Tartagni], Professor of Electronics. We are grateful to our advisors for their time and support!
Our instructors are [http://www.ing2.unibo.it/Ingegneria%20Cesena/Strumenti%20del%20Portale/Cerca/paginaWebDocente?UPN=silvio.cavalcanti@unibo.it Prof. Silvio Cavalcanti], Professor of Biomedical Engineering, Francesca Ceroni, Graduate student in Pharmaceutical Biotechnologies, Sara Montagna, Graduate student in Biomedical Engineering and [http://www-micrel.deis.unibo.it/~christine/ Christine Nardini], PhD in Bioengineering.
Our Project
Introduction
Our goal is the realization of a genetic circuit able to implement the functionality typical of an electronic device called Schmitt Trigger (as defined by its inventor [http://www.otto-schmitt.org/ Dr Otto H. Schmitt]).
Pardon me, what do you mean? (AKA Functional Requirements)
The main characteristics of this device is to be a “smart” switch.
A switch is a device that, given a change in some environmental condition causes an appropriate modification in the output, for instance the output changes from on to off. Often the environmental change is the modification of the value in some parameter (input of the switch) that describes the environment. The device switches for a given value (threshold) of the input. So far so good, however, if the switch input has a value that continually even minimally changes across the threshold, the system will keep going on and off, wasting energy and leaving the system in an unstable state.
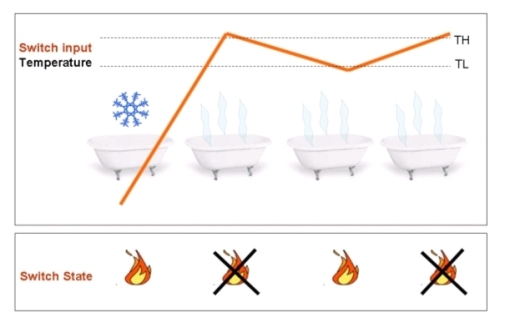
To avoid all this we need a “smart” switch. Basically, this device switches on or off at two different threshold (High and Low threshold, TH and TL respectively) depending on the history of the system. So, according to the state of the system, the threshold for switching will change. This can solve the instability problem, since the variations that cause instability now must be as high as the difference between the two thresholds (see Figure 1).
Figure1. Once the temperature passes the lower threshold, for small changes of the temperature (between TL and TH) the system is able to automatically modify its state (on-off), so that the average temperature of the water remains stable (and so does your mood if you are in the bath tub).
The reasons why the “smart” switch works that well are due to the fact that (1) the system it constitutes has two stable states, that the system reaches when below the lower threshold (TL) or above the higher threshold (TH); and (2) the system has a positive feedback. All this allows for hysteresis to take place, this is the fundamental property we want to reproduce in our genetic circuit.
How? (AKA Technical Requirements)
We wanted to reproduce the same principle with a genetic circuit. To do so we exploited one of such system, existing in the complex of genes that form the Lac Operon shown in Figure 4.
Namely, E.coli can either survive by metabolizing glucose or, in case glucose is missing, lactose. Lactose or glucose metabolism-modes are the two stable state of our system. To perform experiments we use IPTG a structural analog of lactose that cannot be metabolized. Thus the input of our system are the external concentrations of Glucose and IPTG (Gluex and IPTGex respectively).
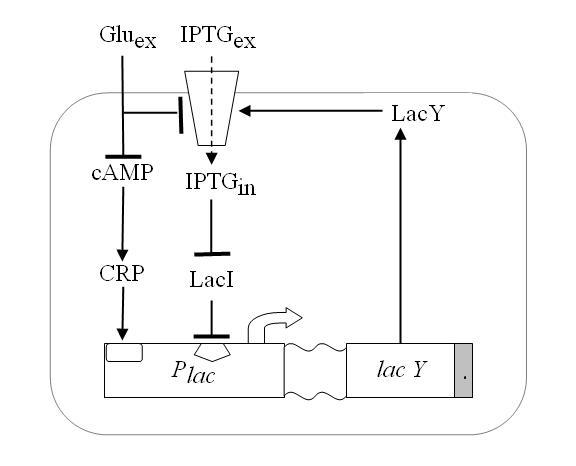
Since lactose metabolism is more energy consuming, usually all the apparatus that takes care of the lactose metabolism is repressed. That is [http://en.wikipedia.org/wiki/Lac_operon the promoter pLac ] is constitutively repressed thank to the presence of the LacI protein that inhibits the transcription of the genes downstream of the operon ([http://bcs.whfreeman.com/mga2e/pages/bcs-main.asp?s=132&n=003&i=283&v=&o=&ns=0&uid=0&rau=0 this] is how it works). When there is IPTG in the environment of the cell, several concurrent processes take place. IPTG enters through the membrane and after some processing it is able to remove the repressor LacI, thus allowing the transcription of genes lacY that in turn enhance the IPTG uptake and increase its action on the repressor LacI. This represents the positive feedback loop we need.
Figure 2. Schematic overview of the Lac operon.
As it is the system can already perform like a Schmitt trigger, for certain values of the inputs. However there is another mechanism that allows the fine tuning of the system: glucose has a double action both on a molecule called cAMP that it bounds to, inhibiting its action on pLac expression (catabolite repression) and on the lactose, since it reduces its uptake (inducer exclusion).
What for?(AKA Applications)
We figured that such a system could be useful for two main purposes.
1. Controlled Gene Expression Trigger
Manipulating gene expression using most of the current protocols in molecular biology usually relay either on a strong overexpression of a protein or on its total silencing. Once induced, both these two extreme perturbations are generally unmodifiable (so that this approaches, although shedding light on the general role of the candidate gene, are rough indicators of the effect of its real physiological transcriptional level). Our genetic device can limit these drawbacks offering 1) a controlled on-off transition of the protein expression at will (by dosing the extracellular IPTG input) and 2) its graded transcriptional level (playing with the extracellular Glucose concentration), representing a powerful tool for biologists. Moreover, in addition to the ability to control the timing and the extent of the induction, our device has the additional advantage to guarantee that small changes in the extracellular concentration of the inducer IPTG would not affect the stability of the switch.
2. Glucose Sensor
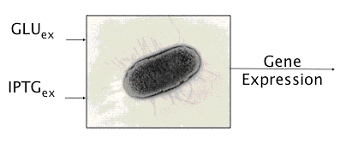
The second application takes advantage of the external glucose dependence. In fact, we know that the expression of the system is stable, but its intensity depends on the glucose. We can reverse the perspective and observe the intensity of the expression to infer the abundance of glucose in the medium.
Figure 3. Schematic view of our device
Mathematical Model
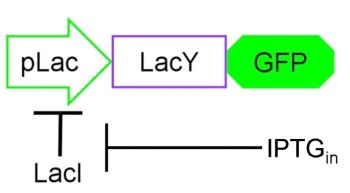
Based on the information we had on the Schmitt Trigger (See in the introducion) and on the [http://en.wikipedia.org/wiki/Lac_operon Lac operon], we decided to explore the behavior of our genetic circuit by mathematically modeling the parts depicted in Figure 4.
Figure 4. The core of our circuit: the pLac promoter, inhibited by protein LacI, and genes LacY and GFP the positive feedback input and reporter gene respectively.
Model
In the following we describe the 3 differential equations we used in the model. They represent the variation over time (derivative d/dt) of the production of the mRNA of lacY (called T for transcript), of the production of the protein LacY (called P for protein) and the variation of the input to our circuit IPTG_in. These equations are sufficient to model the dynamics relevant to the functioning of the system and nor overly complex to avoid this.
Knowing that the β functions represent:
- \beta_D^T glucose dependent LacY transcription inhibitor;
- \beta_R^T IPTG dependent LacY transcription enhancer;
- \beta_D^I glucose dependent IPTG uptake inhibitor;
- \beta_R^I IPTG dependent IPTG uptake enhancer
and have their own formulas and trends.
Since our output is the reporter protein GFP we need another equation to link the protein LacY (P) to GFP, this is simply:
All the parameters are defined here.
To simulate the evolution of these variables we used simulink.
Simulation
With this model, we can predict the circuit behavior and assess if it actually has bistability and hysteresis. In the following we show the results of the simulations.
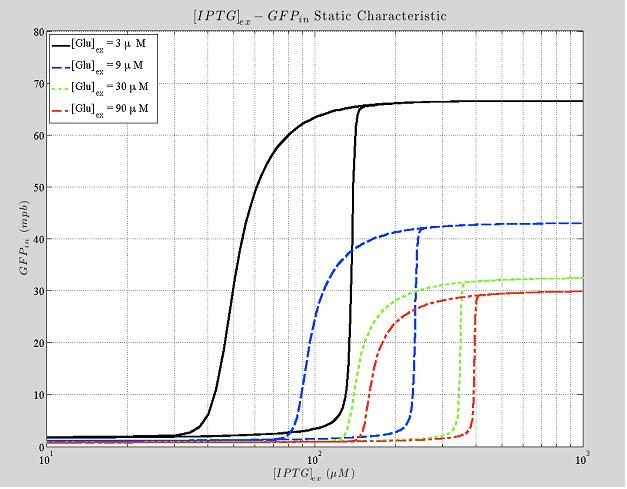
Figure 5. Shows the typical hysteresis profile, different curves are due to different values of the external Glucose.
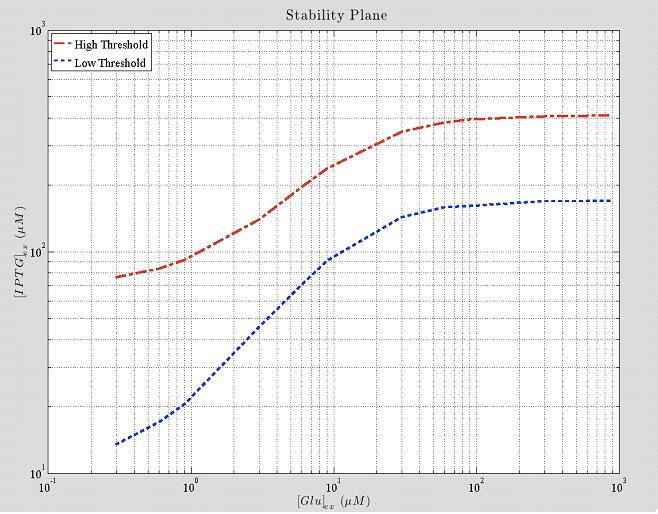
Figure 6. Shows how the 2 thresholds values change for varying levels of external glucose. All the points that lie between the two curves represent unstable states, the system will not remain in that state but it will move to a state in one of the two other areas.
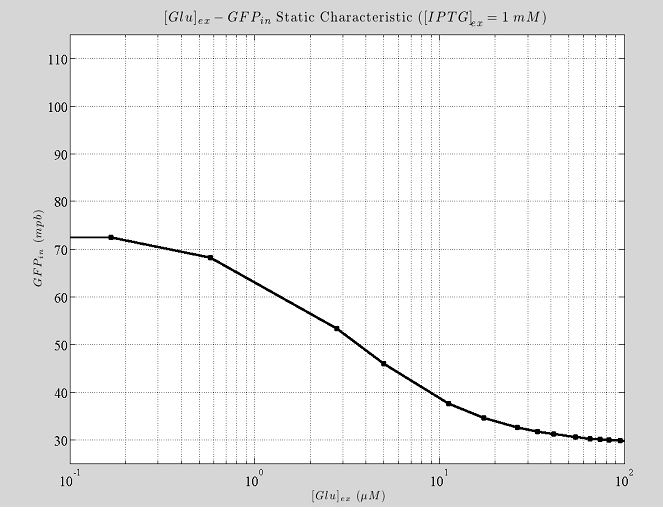
Figure 7.Shows how the fully induced steady state depend on the External Glucose.
Image Acquisition and Elaboration
Experimental SetUp
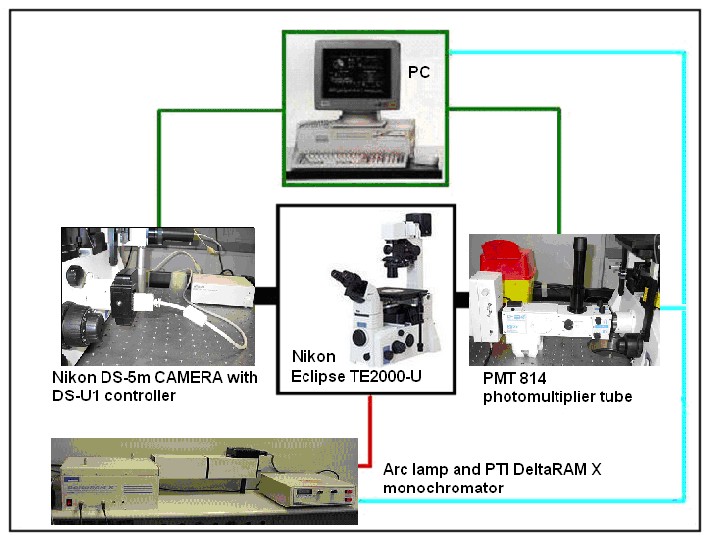
A schematic design of our acquisition system based on the fluorescence microscope is shown in figure.
The illumination system is composed of a 75 Watt Xenon arc lamp connected to a Photon Technology Instruments DeltaRAM X monochromator, which breaks up a single polychromatic light beam into several monochromatic light beams (with only one wavelength each). Only the selected wavelength can pass through the output port and reach the microscope.
The system’s core is a Nikon Eclipse TE2000-U inverted fluorescence microscope. For GFP image acquisition we used a B-2A filter by Nikon with an excitation band between 450 and 490 nm and the optimal emission placed at 520 nm.
The camera used to acquire images and film segments is a Nikon DS-5m with a DS-U1 controller. This one receives the acquired signal form the camera through a serial connection and sends it to the PC through an USB slot. Nikon also supplied an interface software for image acquisition and elaboration.
The Photon Technology Instruments PMT 814 photomultiplier tube is connected to the microscope’s left port, on which the whole light signal can be deviated. Acquisition can be implemented in two different modes, depending on the signal amplitude.
The control software is implemented in a Labview environment and permits the regulation of the excitation wavelength and the calibration of the system. It also pictures the output signal from the photomultiplier, which can be memorized and elaborated.
Image Acquisition
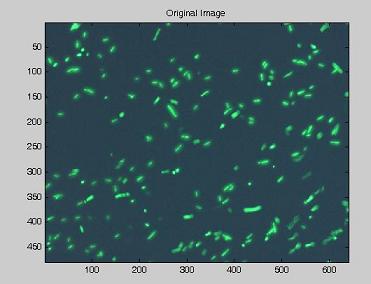
Using the experimental set up illustrated here, we obtain short movies of fluorescent bacteria; then we extract several frames from each of them, using a dedicated software. Example images are shown below.
Image Elaboration
Since we need to know how much of the total image area is occupied by bacteria, we process the images with our segmentation algorithm implemented in Matlab.
Here are the processing phases:
- the RGB image is read and the green channel is extracted;
- a morphological top hat filtering is performed on the grayscale image;
- by means of an adaptive threshold, the image is tranformed in a binary one;
- the distance between every point and the nearest black pixel is calculated;
- on this last image the [http://en.wikipedia.org/wiki/Watershed_%28algorithm%29 watershed] algorithm is applied: every pixel is assigned a label, depending on the segmentated region it belongs; then, every labeled region is represented with a different colour, as in the images shown below;
- the area of every segmented region is calculated, checking that the summation of these areas (including the background) balances the image total area;
- two complementary matrices are created:
-ImageWithOutBackground, containing the intensity positive values corresponding only to pixels recognised as bacteria, with zeros elsewhere; -ImageBackground, which contains the intensity positive values of pixels recognised as background, with zeros as other entries;
- with a summation over all the ImageWithOutBackground matrix' entries, the total intensity is obtained;
- dividing this value by the total bacteria area, the output is the normalized intensity we use to compare the fluorescence of different kinds of bacteria.
Thanks to Camilo Melani for his competence and kindness during the algorithm implementation.
Biodevice
Components
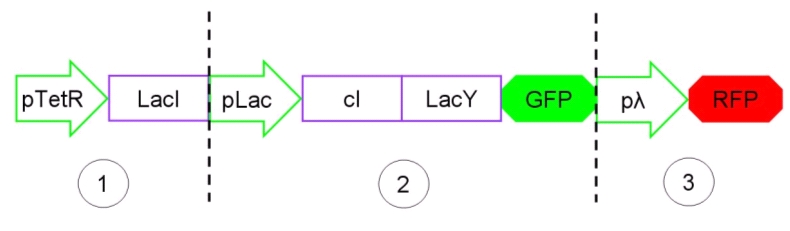
The biodevice Genetic Schmitt Trigger [http://partsregistry.org/Part:BBa_I763029 (I763029)] built with IGEM 2007 Biobricks consists of 3 main parts combined in the same plasmid: pTeTR-LacI [http://partsregistry.org/Part:BBa_I763026 (I763026)], pLac-cI-LacY-GFP [http://partsregistry.org/Part:BBa_I763019 (I763019)], pλ-RFP [http://partsregistry.org/Part:BBa_I763007 (I763007)].
Each part displays a specific function depending on the promoters and coded genes.
Figure 1: The Genetic Schmitt Trigger biodevice ([http://partsregistry.org/Part:BBa_I763029 I763029]).
- 1.pTeTR-LacI [http://partsregistry.org/Part:BBa_I763026 (I763026)] codes for LacI gene [http://partsregistry.org/Part:BBa_C0012 (C0012)] regulated by pTeTR [http://partsregistry.org/Part:BBa_R0040 (R0040)] inverting promoter. pTeTR can be consider a constitutive promoter in absence of tetracycline or its analog, aTc. Its action is inhibited by the addition of this antibiotic. This promoter regulates the expression of LacI gene whose protein inhibits the activation of pLac promoter. This part is important to make up for endogenous LacI and to prevent pLac activation in absence of induction with lactose or IPTG.
- 2.pLac-cI-LacY-GFP [http://partsregistry.org/Part:BBa_I763019 (I763019)]: LacY permease [http://partsregistry.org/Part:BBa_J22101 (J22101)] controlled by pLac promoter [http://partsregistry.org/Part:BBa_R0010 (R0010)] introduces a positive feedback necessary for hysteresis. LacY is a lactose membrane transporter which allows the uptake of Lactose (or IPTG) in the cell. Lactose (or IPTG) on his own, causes the LacI repressor release from pLac operators. At the same time GFP proteins [http://partsregistry.org/Part:BBa_J04031 (J04031)] are produced as reporters of the induction. We also introduce the gene for cI repressor [http://partsregistry.org/Part:BBa_C0051 (C0051)] that binds to the cI regulator [http://partsregistry.org/Part:BBa_R0051 (R0051)] and inhibits its action.
- 3.pλ-RFP [http://partsregistry.org/Part:BBa_I763007 (I763007)] acts as reporter of standard condition. Pλ promoter [http://partsregistry.org/Part:BBa_R0051 (R0051)] is a constitutive promoter from phage-λ. It regulates RFP protein [http://partsregistry.org/Part:BBa_E1010 (E1010)] expression and it is repressed by cI repressor [http://partsregistry.org/Part:BBa_C0051 (C0051)]. So, when external glucose is high, pLac transcription level, cI and GFP production are high, while RFP production is low; when external glucose is low pLac transcription level, cI and GFP production are low while RFP production is high. As a consequence, we can consider RFP fluorescence levels as an indicator of glucose concentration in culture medium.
How it works
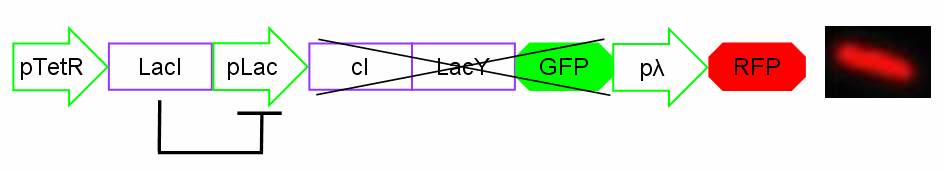
In standard conditions, without IPTG induction pLac promoter is repressed by the LacI repressor binding on the operator sites and cI, LacY and GFP proteins are not expressed. In this condition pλ promoter is active and RFP is expressed. The cells show red fluorescence.
Figure 2: Project in action: no IPTG in the medium.
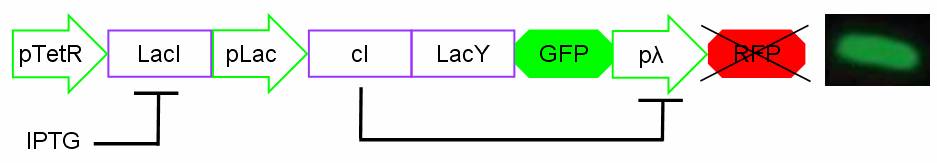
For induction IPTG is added in the culture medium. Due to inductor binding to LacI repressor, Plac promoter is activated and cI, LacY and GFP genes are expressed. In this condition pλ promoter is repressed by cI inhibitor and so RFP is not expressed. The cells show green fluorescence.
Figure 3: Project in action: IPTG in the medium.
Intermediates
The table below shows all the intermediates that we have obtained to build up our final device. For each part you can find: -in the first column the link to the Registry;
-in the second column the components for each part;
-in the third column the plasmid the part has been cloned in;
-in the fourth the insert length after double digestion with Xba1/Pst1 enzymes linked to the electrophoresis gel run;
-in the fifth column cell vitality after transformation;
-in the sixth column fluo data for each part.
| Igem Code | Fragment | Plasmid | Length (bp) | Vitality | Fluo Data | |
|
|
|
|
|
Clicking on the length of the fragment you will be shown the electrophoresis gel of the digestion of the fragment excised from the plasmid by cutting it with a double digestion using Xba and PstI enzymes.
Acknowledgments
Our Team was funded by:
- European Union SYNBIOCOMM project

- Ser.In.Ar. Cesena
- University of Bologna - Cesena Campus
Overview Table
| Our Team | Project Design | Project Results |
|
|